Passionate about gene expression regulation, I performed my PhD in epigenetics at Institut Pasteur. During my PhD, I contributed to the discovery of a new mechanism of alternative splicing regulation, involving histone protein post-translational modifications (Lavigne et al., PLoS Genetics 2009; Saint-André et al., Nature Structural and Molecular Biology 2011).
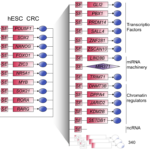
After that, I added to my experience of experimental molecular biology, an experience in computational biology, through working for 3 years as a post-doctoral associate of the Whitehead Institute for Biomedical Research of MIT. There, I contributed to the study of “super-enhancers”, and developed a method and a bioinformatic tool to predict the transcription factors at the core of each cell type or cell state’s gene expression program (Hnisz, Abraham and Lee et al., Cell 2013; Tan et al., Molecular Cell 2016; Saint-André et al., Genome Research 2016; Saint-André et al., US Patent 2022).
I then performed a 2-year post-doc at the Curie Institute, where I could identify a long non-coding RNA signature that separates sub-populations of immune cells upon diversification and contributed to evidence upstream control of this regulation by a TNF communication loop (Alculumbre et al., Nature Immunology 2018).
In 2018, I joined the Bioinformatics and Biostatistics HUB, where I am detached to work on the “Milieu Intérieur” project with the Translational Immunology Unit. Our aim is to better understand human immune response determinants and participate to precision medicine development. I yet contributed to multiple works on “Milieu Intérieur” data (Possémé et al., Cell Reports 2022 ; Delavy et al., Gut Microbes 2023), and studies related to tuberculosis (Libre et al., Frontiers in Immunology 2022), HIV (Lista et al., Retrovirology 2023) and Covid-19 (Smith, Possémé, Bondet et al., Nature communications 2022).
Through multi-variate data analysis of Milieu Intérieur (MI) proteomic data and integration with genomic, epigenomic, cellular, clinical, environmental, and nutritional factors related to the donors of the MI cohort, we recently found that age, sex, smoking status, cytomegalovirus (CMV) serostatus and body mass index (BMI) are significant drivers of immune response variability. Smoking effects explained up to 9% of the inter-individual variability of specific cytokine responses, a level equivalent to the effects of age, sex or common genetic variants. We observed that active smoking modifies cytokine responses to innate and adaptive immune stimulations, with persistent effects on adaptive immunity. We showed that this persistent effect is mediated through specific cell subsets and DNA methylation on signal transducers and metabolism regulators, supporting a long-term epigenetic memory of the past smoking status of individuals (Saint-André et al., 2024). This work made the cover of the 8000th issue of Nature, had a Nature “news” and “news and views” companion articles and and generated multiple press articles in large audience journals (Le Monde, El Pais, Liberation, Pour la Science, Courier International, The Scientist, US News, Scientific American,…) , 4 radio (France Inter 19h16 24/02/14, France Info, France Culture, Radio Canada) and 3 TV interventions (BFM TV, France TV Info, CNN).
I also pursue my interest in the development of bioinformatic methods to study transcriptional regulatory networks (Saint-André et al., Computational and Structural Biotechnology Journal 2021), that we are now applying to “Milieu Intérieur” transcriptomic data.
In parallel of my research work, I have set up or contribute to the following courses:
– “ChIP-seq data analysis” (1 day per session), in “Bioinformatics program for PhD students”, Institut Pasteur, Paris, since 2019
– “Multivariate Data Analysis” (2 days per session, in french or english), in “R Programming and Statistics”, Institut Pasteur, Paris, since 2018
– “IA et Santé” course (one-week), Centrale-Supelec, Paris, since 2019
– Invited lectures, Ecole Centrale de Nantes, since 2019
I am also regularly training students and participated in the following conference organization or evaluation committees:
– Member of the organization committee of JOBIM, Paris 2021
– Invited member for oral and poster evaluation at the YRLS conference, Paris 2019 and 2023
– Jury member for Societé Française de Bioinformatique (SFBI) poster price, JOBIM 2019