The proteomics facility, headed by Mariette Matondo, is part of the Mass Spectrometry for Biology Utechs (MSBio) directed by Julia Chamot-Rooke. The overall objective of the platform is to develop innovative bottom-up proteomics strategies to address protein function, regulation and dynamic interactions, in particular in the context of infectious diseases. Our collaborators benefit from the expertise of the staff, as well as state-of-the-art instrumentation, including the latest generation of high speed, high-resolution mass spectrometers (Orbitrap), and bioinformatics tools. The proteomics analyses provided by the platform include:
- Identification of low abundant proteins in complex biological matrices
- Advanced quantitative proteomic strategies
- Characterization of post-translational modifications (PTMs)
- Analysis and dynamics of protein complexes
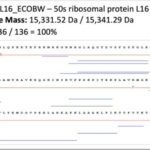
- Top-Down Proteomics to identify proteoforms
Analyses can be performed either as research collaboration or service. Prior to starting the project, an experimental design is defined and the staff provides advice on sample preparation. Results are further provided after a bioinformatic analysis of the data.
Please contact Mariette Matondo : mariette.matondo@pasteur.fr for more information
User Committee members : Marco Bellinzoni, Serge Bonnefoy, Christel Brou, Alexandre Chenal, Ludovic Deriano , Olivier Dussurget , Simonetta Gribaldo, Guilhem Janbon,Pierre Alexandr Kaminski, Pierre-Lê Bury, Chloé Lescale ,Pascale Pescher , Cosmin Saveanu, Timothy Wai
Steering committee members : Sandrine SAGAN (CNRS), Cosmin Saveanu, Joëlle VINH (ESPCI)
How to work with us / How to apply for support
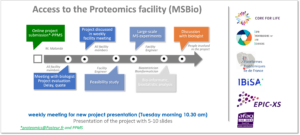
For project proposals, please submit your project request here on PPMS (stratocore): for the facility use MSBio Core Facilty Request Form : https://ppms.eu/pasteur/req/?pf=29&project=true&form=68. Additionally, you can complete this form and send it to proteomics@pasteur.fr.
If you have any suggestion on how we can improve our service, please do not hesitate to contact us: Reclamation / Suggestions
Open Desk
On Tuesday at 10.30 am. Please contact Mariette Matondo at proteomics@pasteur.fr
Satisfaction Surveys
The following files link to the analysis of the surveys
- 2017 : 2017 LimeSurvey-Result
- 2018 : 2018-LimeSurvey-Result
- 2019 : 2019-LimeSurvey-Result
- 2020 : 2020_Satisfaction survey
- 2021 : 2021_Satisfaction survey
- 2022 : 2022_Satisfaction survey
The proteomic faciltiy is certified ISO 9001
The proteomics facility is IBisA labelled
Practical Information
We are part of the C2RT (Center for Technological Resources and Research), a center gathering the Technology and Service units (UTechS) and the technological core facilities of the campus. As such we follow the common guidelines and good practices
If you did not find what you were looking for, please check out the DT Brochure