Protein folding and membrane interactions
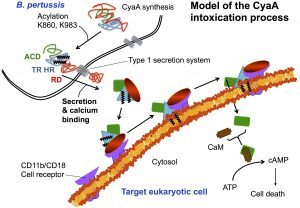
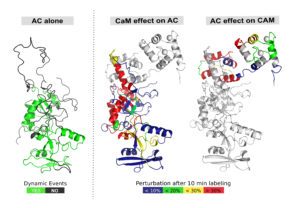
My research interests are mainly focused on protein folding, thermodynamics, hydrodynamics and protein translocation across membranes. In the BIM Unit, I am characterizing a bacterial toxin, the adenylate cyclase (CyaA) produced by Bordetella pertussis, the causative agent of whooping cough, which is currently in increasing incidence and represents a global public health concern. The study of CyaA offers a unique opportunity to explore various topics such as intrinsically disordered proteins (IDP), the effect of molecular crowding, protein-protein, protein-ligand and protein-membrane interactions. More about the CyaA project can be found on the page of the Group.
Besides investigating the biophysics of CyaA, I have pursued several projects initiated during my previous post-doctoral positions or within new collaborations established with various groups inside or outside Institut Pasteur. The main goal is to improve the understanding of the behavior of amphitropic proteins, i.e., to describe how soluble proteins are able to partition from the solution to the membrane (and eventually translocate across the lipid bilayer) to regulate their functions. The methods used to characterize biophysics of amphitropic proteins are mainly biochemical and spectroscopic approaches. Inside Institut Pasteur, I am working with other Units from Institut Pasteur on these topics, like PimA with Pedro Alzari’ Unit and large clostridial toxins with Michel Popoff’ Unit. Outside IP, I am collaborating with several groups, such as WJ Tang, Chicago, USA, Marcelo Guerin, Bilbao, Spain, Scot Ouellette, South Dakota, USA, on protein membrane interactions; Abdel Aouacheria, Lyon and Vincent Forge, Grenoble on Bcl proteins; Allen Minton, Bethesda, USA, on molecular crowding; Daniel Isabey, Creteil Mondor, on multi-scale study of CyaA effects on the upper respiratory tract, …
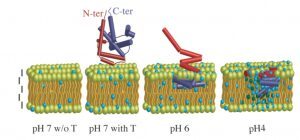
Illustration of the pH-dependent membrane insertion of the translocation domain of the diphtheria toxin, inferred from specular neutron reflectometry, solid state NMR and other biophysical methods (figure from Chenal et al., 2009, J. Mol. Biol.).
A: X-ray Structure of Anthrolysin O, a 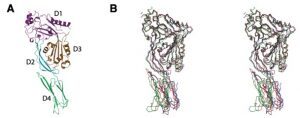
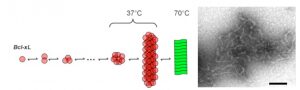
Schematic view of Bcl-XL amyloidogenesis and EM of fibrils (figure from Chenal et al., 2012, J. Mol. Biol.).