I received my Ph.D. in Molecular Chemistry from the Ecole Polytechnique (Palaiseau, France) in 2010 under the supervision of Julia Chamot-Rooke. During this time, I was involved in the development of a top-down mass spectrometry approach combined with hydrogen/deuterium exchange (HDX) for the structural analysis of protein complexes. After my PhD, I joined the proteomics core facility of the Institut Pasteur to work on the Orbitrap Velos ETD coupled with a liquid chromatography. I also manage the high-throughput bottom-up proteomics analysis. I divide my time between the core facility and the Structural Mass Spectrometry and Proteomics Unit headed by Julia Chamot-Rooke, where I am involved in the development of top-down proteomics using high resolution mass spectrometers (Orbitraps).
Click to view graph
Connections
Click to view timeline
Timeline
Projects
Software
Publications
Download-
2023The cell wall lipoprotein CD1687 acts as a DNA binding protein during deoxycholate-induced biofilm formation in Clostridioides difficile., NPJ Biofilms Microbiomes 2023 May; 9(1): 24.
-
2023Hemisynthetic alkaloids derived from trilobine are antimalarials with sustained activity in multidrug-resistant Plasmodium falciparum., iScience 2023 Feb; 26(2): 105940.
-
2023Widespread amyloidogenicity potential of multiple myeloma patient-derived immunoglobulin light chains., BMC Biol 2023 Feb; 21(1): 21.
-
2022Bacterial Membrane Vesicles as a Novel Strategy for Extrusion of Antimicrobial Bismuth Drug in Helicobacter pylori., mBio 2022 Oct; 13(5): e0163322.
-
2022A time-resolved multi-omics atlas of Acanthamoeba castellanii encystment., Nat Commun 2022 Jul; 13(1): 4104.
-
2022NAD kinase promotes Staphylococcus aureus pathogenesis by supporting production of virulence factors and protective enzymes., Elife 2022 Jun; 11(): .
-
2022A novel screening strategy to identify histone methyltransferase inhibitors reveals a crosstalk between DOT1L and CARM1., RSC Chem Biol 2022 Apr; 3(4): 456-467.
-
2021Purification of infection-associated macropinosomes by magnetic isolation for proteomic characterization, Nature Protocols, 2021, 16 (11), pp.5220-5249. ⟨10.1038/s41596-021-00610-5⟩.
-
2021De Novo Sequencing of Antibody Light Chain Proteoforms from Patients with Multiple Myeloma., Anal Chem 2021 08; 93(30): 10627-10634.
-
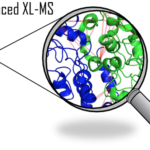
2021Advanced In Vivo Cross-Linking Mass Spectrometry Platform to Characterize Proteome-Wide Protein Interactions., Anal Chem 2021 Feb; (): .
-
+View full list of publications