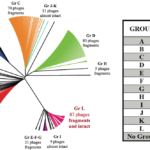
Click to view graph
Connections
Click to view timeline
Timeline
Courses
Workshop: Bacterial RNAseq
We propose a skills training for PhD students on Bacterial RNAseq experiments Using total RNAs, we will i) enrich the mRNAs by depleting the rRNAs ; ii) prepare and sequence cDNA library and iii) analyse […]
2016-06-06 09:00:00
2016-06-10 17:00:00
Europe/Paris
Workshop: Bacterial RNAseq
We propose a skills training for PhD students on Bacterial RNAseq experiments Using total RNAs, we will i) enrich the mRNAs by depleting the rRNAs ; ii) prepare and sequence cDNA library and iii) analyse […]
28 Rue du Docteur Roux, Paris, France
Transversal Projects
Projects
Software
Publications
Download-
2024Spores of Clostridioides difficile are toxin delivery vehicles., Commun Biol 2024 Jul; 7(1): 839.
-
2024A monoclonal antibody collection for C. difficile typing ?, Gut Pathog 2024 Jan; 16(1): 4.
-
2024Anti-S-layer monoclonal antibodies impact Clostridioides difficile physiology., Gut Microbes 2024 ; 16(1): 2301147.
-
2024SpoIVA is an essential morphogenetic protein for the formation of heat- and lysozyme-resistant spores in Clostridium sporogenes NBRC 14293., Front Microbiol 2024 ; 15(): 1338751.
-
2023Extracellular succinate induces spatially organized biofilm formation in Clostridioides difficile., Biofilm 2023 Dec; 5(): 100125.
-
2023Regulation of Clostridial Toxin Gene Expression: A Pasteurian Tradition., Toxins (Basel) 2023 Jun; 15(7): .
-
2023The cell wall lipoprotein CD1687 acts as a DNA binding protein during deoxycholate-induced biofilm formation in Clostridioides difficile., NPJ Biofilms Microbiomes 2023 May; 9(1): 24.
-
2023Elucidating dynamic anaerobe metabolism with HRMAS 13C NMR and genome-scale modeling., Nat Chem Biol 2023 May; 19(5): 556-564.
-
2022c-di-AMP signaling is required for bile salt resistance, osmotolerance, and long-term host colonization by Clostridioides difficile., Sci Signal 2022 Sep; 15(750): eabn8171.
-
2022Core-, pan- and accessory genome analyses of Clostridium neonatale: insights into genetic diversity., Microb Genom 2022 May; 8(5): .
-
+View full list of publications