After a PhD in bioinformatics and microbial ecology, I joined the team of C. Brochier-Armanet (Laboratoire de Biométrie et Biologie Évolutive) for a 2 years postdoc to study the deep evolution history of Archaea and Bacteria. Then I performed a postdoc at Institut Pasteur in S Gribaldo’s team to develop phylogenomic approaches aiming at tackling ancient evolutionary events within the microbial world. Since January 2018, I integrated the HUB as a permanent bioinformatician and I am involved in phylogenomic, metagenomics and comparative genomics projects.
Courses
PHINDaccess – Comparative Genomics
Comparative genomics is a field of biological research in which a variety of tools are used to compare genome sequences of different species. By carefully comparing characteristics that define various organisms, researchers can pinpoint […]
Introduction à la Phylogénie Moléculaire : Concepts, méthodes et interprétation
Ce cours a pour objectif d’enseigner les bases théoriques et pratiques nécessaires à la conduite d’analyses phylogénétiques. Les sessions théoriques seront consacrées aux bases du domaine (représentation et combinatoire des arbres, modèles d’évolution) et […]
Introduction to Molecular Phylogenetics – ENS Paris
Introduction to Molecular Phylogenetics course aims to give the basic theoretical and practical concepts, best practices, and software necessary to start working on molecular phylogenetics and its applications to epidemiology. The course will have […]
Introduction à la Phylogénie Moléculaire : Concepts, méthodes et interprétation
Ce cours a pour objectif de donner les bases théoriques et pratiques, ainsi que les bonnes pratiques nécessaires au démarrage de travaux en phylogénie et épidémiologie moléculaires. Les sessions théoriques donneront les bases du […]
Projects
CV
Experiences
-
2018Bioinformatician
Institut Pasteur (EBMC Unit – Bioinformatics and Biostatistics HUB)2017Postdoctoral fellow (9 months)
Institut Pasteur (BMGE Unit)2015Postdoctoral fellow (2 years)
Laboratoire de Biométrie et Biologie Évolutive, Université Lyon 12013Postdoctoral fellow (1,5 years)
Laboratoire Microorganismes : Génome et Environnement, Université Clermont
Education
-
2013Phd in Bioinformatics at Université Blaise Pascal, Clermont-Ferrand
Exploring the microbial diversity from high throughput sequencing2009Master 2 research at Université Blaise Pascal, Clermont-Ferrand
Analysis and modelling of biological data2007Diploma in Agronomic engineering from Institut Agronomique et Vétérinaire Hassan II
Rabat – Morocco
Publications
Download-
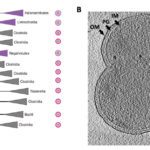
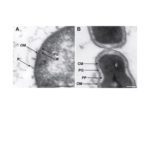
2025Terrabacteria: redefining bacterial envelope diversity, biogenesis and evolution., Nat Rev Microbiol 2025 Jan; 23(1): 41-56.
-
2024Borg extrachromosomal elements of methane-oxidizing archaea have conserved and expressed genetic repertoires, Nature Communications, 2024, 15 (1), pp.5414. ⟨10.1038/s41467-024-49548-8⟩.
-
2024Proteins containing photosynthetic reaction centre domains modulate FtsZ-based archaeal cell division, Nature Microbiology, 2024, 9 (3), pp.698-711. ⟨10.1038/s41564-024-01600-5⟩.
-
2023Diversification of division mechanisms in endospore-forming bacteria revealed by analyses of peptidoglycan synthesis in Clostridioides difficile., Nat Commun 2023 Dec; 14(1): 7975.
-
2023The Mla system of diderm Firmicute Veillonella parvula reveals an ancestral transenvelope bridge for phospholipid trafficking., Nat Commun 2023 Nov; 14(1): 7642.
-
2022Ancient origin and constrained evolution of the division and cell wall gene cluster in Bacteria., Nat Microbiol 2022 Dec; 7(12): 2114-2127.
-
2022Genetic Screens Identify Additional Genes Implicated in Envelope Remodeling during the Engulfment Stage of Bacillus subtilis Sporulation., mBio 2022 Oct; 13(5): e0173222.
-
2022Localization and functional characterization of the alternative oxidase in Naegleria., J Eukaryot Microbiol 2022 Jul; 69(4): e12908.
-
2022An ancient divide in outer membrane tethering systems in bacteria suggests a mechanism for the diderm-to-monoderm transition., Nat Microbiol 2022 Mar; 7(3): 411-422.
-
2021Comparative genomic analysis of Methanimicrococcus blatticola provides insights into host adaptation in archaea and the evolution of methanogenesis, Thomas, C.M., Taib, N., Gribaldo, S. et al. Comparative genomic analysis of Methanimicrococcus blatticola provides insights into host adaptation in archaea and the evolution of methanogenesis. ISME COMMUN. 1, 47 (2021)..
-
+View full list of publications