I work as Research Software Engineer in the Bioinformatics and Biostatistics HUB of Institut Pasteur. My main mission is to help researchers to create the most efficient, scalable, and sustainable research codes possible in order to enable new scientific advances.
Courses
Analyse de données reproductible
La production de données de plus en plus massives, le développement de méthodes d’analyse de plus en plus complexes, et le nombre en constante augmentation d’outils disponibles amènent les scientifiques à développer des chaines de traitement (workflows) de plus […]
Scientific Programming in Python
The ever growing usage of high throughput technologies in Biology is revolutionizing the life sciences and profoundly changing its practices. Scripting languages are used on a daily basis in life science labs in order […]
Introduction to Python Programming
Python is often used to learn programming as it is very readable and easy to write. Python is a very popular language in bioinformatics where many programs and libraries are written in python. This […]
Introduction to data analysis
This five weeks graduate level course will give participants basic skills and hands-on training in biostatistics. It will cover all the steps of an analysis workflow : design, collection, curation, hypothesis testing and data […]
Projects
Software
Tools
Former Teams
CV
Activities
- Help scientists to develop new tools (architecture, design, implementation).
- O|B|F (http://www.open-bio.org/) member.
Skills
- Strong programming experience in Advanced Python.
- Software architecture and design.
- NoSQL DataBase (MongoDB, CouchDB)
- XML/YAML
- continuous integration (github/travis-CI/readthedocs, gitlab/gitlab-CI)
- containers (Docker, Apptainer/Singularity)
- linux (Gentoo, Xubuntu)
Main projects on the campus
- MacsyFinder (ongoing project)
- IntegronFinder (ongoing project)
- correlationplus (former project)
- bioconvert (former project)
- CRAW (former project)
- Mobyle (former project)
- http://Mobyle.pasteur.fr
- Mobyle: a new full web bioinformatics framework

Teaching
Education
- 2002 Phd in Molecular and cellular biology.
- “Rôle de deux protéines QN1 et PATF impliquées dans l’arrêt de prolifération des cellules de la neurorétine aviaire au cours du developpement”.
- 2001 “Informatique En Biologie” course (Pasteur)
Publications
Download-
2024Integron cassettes integrate into bacterial genomes via widespread non-classical attG sites., Nat Microbiol 2024 Jan; 9(1): 228-240.
-
2023BioConvert: a comprehensive format converter for life sciences., NAR Genom Bioinform 2023 Sep; 5(3): lqad074.
-
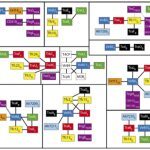
2023MacSyFinder v2: Improved modelling and search engine to identify molecular systems in genomes, Peer Community Journal, 2023, 3, pp.e28. ⟨10.24072/pcjournal.250⟩.
-
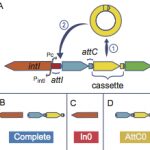
2022IntegronFinder 2.0: Identification and Analysis of Integrons across Bacteria, with a Focus on Antibiotic Resistance in Klebsiella., Microorganisms 2022 Mar; 10(4): .
-
2021Extracting Dynamical Correlations and Identifying Key Residues for Allosteric Communication in Proteins by correlationplus., J Chem Inf Model 2021 Oct; 61(10): 4832-4838.
-
2018CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins, Nucleic Acids Res. 2018 Jul;46(W1):W246-W251.
-
2018Antisense transcriptional interference mediates condition-specific gene repression in budding yeast, Nucleic Acids Res. 2018 May;.
-
2017Abundance and co-occurrence of extracellular capsules increase environmental breadth: Implications for the emergence of pathogens, PLoS Pathog. 2017 Jul;13(7):e1006525.
-
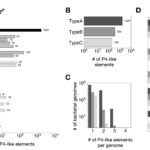
2016Identification and analysis of integrons and cassette arrays in bacterial genomes., Nucleic Acids Res. 2016 Apr 29. pii: gkw319..
-
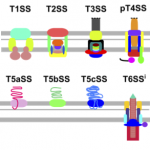
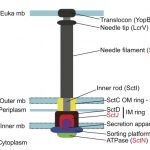
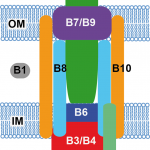
2016Identification of protein secretion systems in bacterial genomes, Scientific Reports 6, 23080 (2016).
-
+View full list of publications


 https://orcid.org/0000-0002-0220-0482
https://orcid.org/0000-0002-0220-0482