I joined the Bioinformatics and Biostatistics Hub at Institut Pasteur in 2016 and was detached to the Platform Biomics (http://biomics.pasteur.fr). Since Oct 2018, I am leading the SALSA group (a.k.a. dry-lab group) activities, which is currently composed of Laure Lemée, Etienne Kornobis, Rania Ouazahrou and myself. Dimitri Desvillechabrol contributed to the group with numerous software development from March 2021 to May 2023. Rachel Legendre and Hugo Varet were also present from Sept 2018 to Nov 2019). We also host one master per year on average as well as visits from the international Pasteur Network.
I have an interdisciplinary research experience: after a PhD in Astronomy (gravitational wave data analysis), I joined several research institutes to work in the fields of plant modelling (INRIA, Montpellier, 2008-2011), System Biology — in particular logical modelling (EMBL-EBI Cambridge, U.K., 2011-2015), and drug discovery (Sanger Institute, Cambridge, U.K.), 2015). On a daily basis, I use data analysis and machine learning techniques within high-quality software to tackle scientific problems. Examples of scientific software are bioservices (access to biological web services programmatically), Sequana (Pipelines related to NGS analysis including genomics, transcriptomics, short and long reads), Bioconvert (a collaborative project to convert biological formats), Damona and a set of NGS pipelines including RNA-seq, ChiP-seq, denovo, variant calling, long reads assemblies, etc…
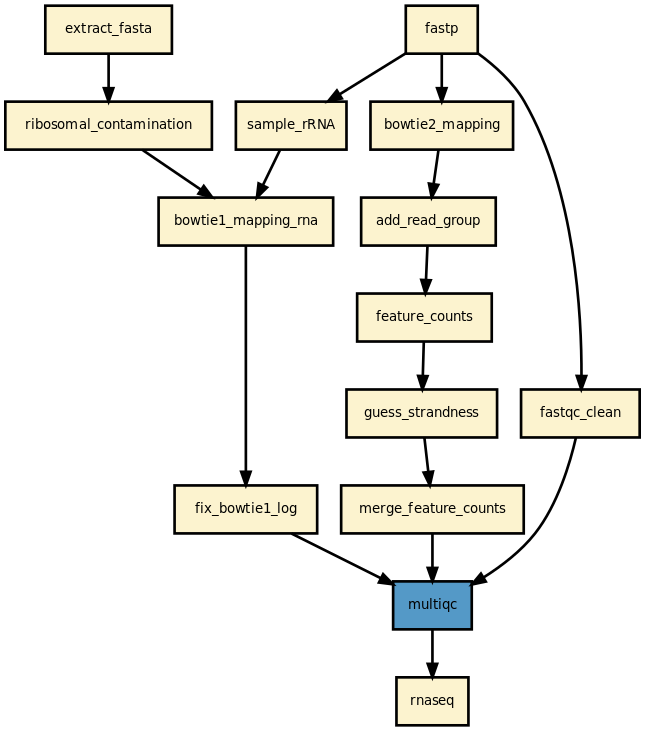 Sequana is a versatile tool that provides (i) a Python library dedicated to NGS analysis (e.g., tools to visualise standard NGS formats), (ii) a set of pipelines dedicated to NGS in the form of Snakefiles (Makefile-like with Python syntax based on snakemake framework), (iii) original tools to help in the creation of such pipelines including HTML reports. Examples of standalone applications are
Sequana is a versatile tool that provides (i) a Python library dedicated to NGS analysis (e.g., tools to visualise standard NGS formats), (ii) a set of pipelines dedicated to NGS in the form of Snakefiles (Makefile-like with Python syntax based on snakemake framework), (iii) original tools to help in the creation of such pipelines including HTML reports. Examples of standalone applications are
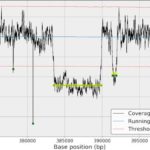
- sequana_coverage ease the extraction of genomic regions of interest and genome coverage information
- sequana_taxonomy performs a quick taxonomy of your FastQ. This requires dedicated databases to be downloaded.
- Sequanix: GUI for snakemake workflows, a GUI for Snakemake workflows (hence Sequana pipelines as well)
Sequana can be used by developers to create new pipelines and by users in the form of applications ready for production.
Would you be interested in joining our software development efforts, or interested in providing NGS snakemake pipelines, please have a look at Sequana or simply contact me.
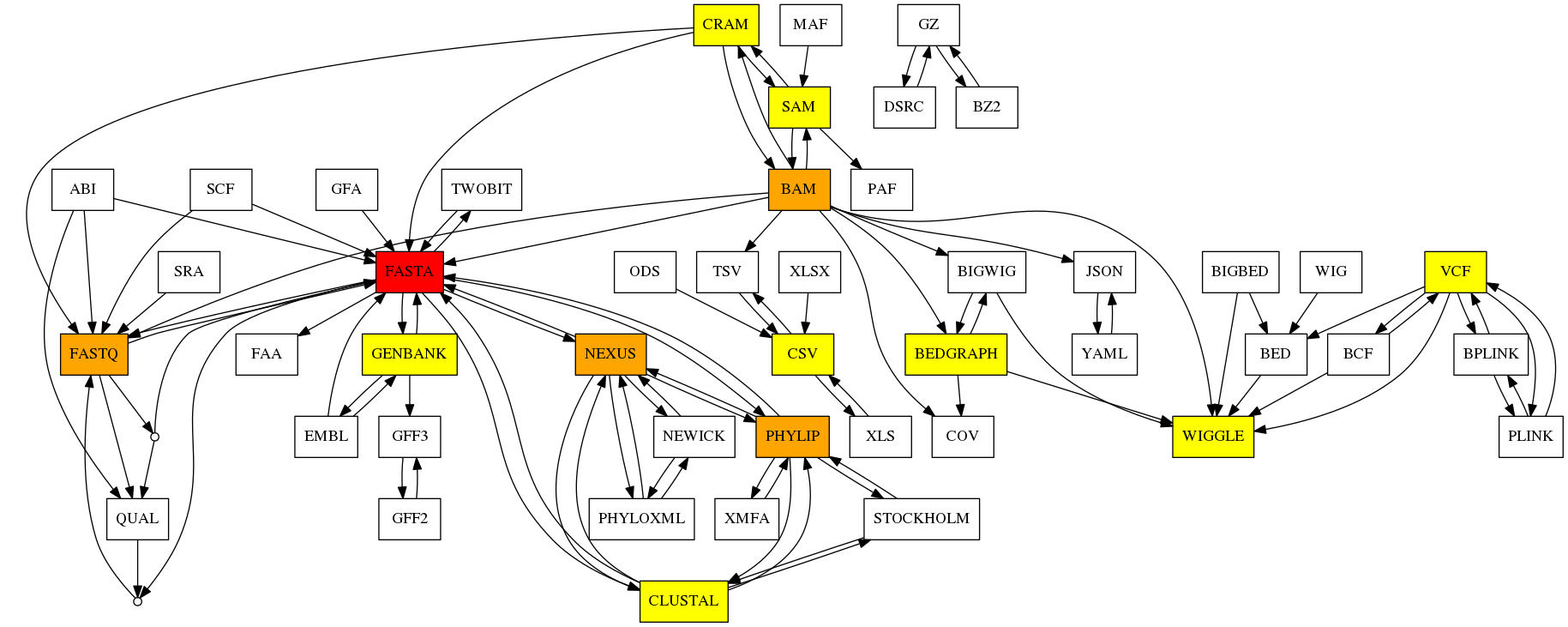
The other on-going project is Bioconvert a collaborative project where everybody is welcome to contribute with code, ideas, examples, documentation, etc.