After an initial degree in Biology, I started my professional career as a software developer. Following the completion of my software engineering degree at the CNAM, I joined the Institut Pasteur, where I initially worked in the “Software and Databases” group. I joined the Bioinformatics and Biostatistics Hub within the Center of Bioinformatics, Biostatistics and Integrative Biology (C3BI) as a Research Engineer in 2017, where I lead the WINTER group.
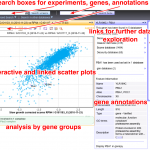
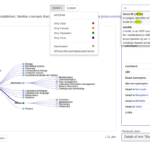
Click to view graph
Connections
Click to view timeline
Timeline
Courses
Software Carpentry
Software Carpentry aims to help researchers get their work done in less time and with less pain by teaching them basic research computing skills. This hands-on workshop will cover basic concepts and tools, including […]
2018-03-28 09:00:00
2018-03-29 17:00:00
Europe/Paris
Software Carpentry
Software Carpentry aims to help researchers get their work done in less time and with less pain by teaching them basic research computing skills. This hands-on workshop will cover basic concepts and tools, including […]
Pasteur, Paris, France
Projects
Software
Tools
Former Teams
Publications
Download-
2022Methods Included, Communications of the ACM, 2022.
-
2022HARIBOSS: a curated database of RNA-small molecules structures to aid rational drug design, bioRxiv (2022) 10.1101/2022.05.17.492306.
-
2021ELIXIR and Toxicology: a community in development, F1000Research 2021, 10(ELIXIR):1129.
-
2021Multitrait GWAS to connect disease variants and biological mechanisms., PLoS Genet 2021 Aug; 17(8): e1009713.
-
2021Gene expression analysis in EBV-infected ataxia-telangiectasia cell lines by RNA-sequencing reveals protein synthesis defect and immune abnormalities., Orphanet J Rare Dis 2021 06; 16(1): 288.
-
2021Viral Host Range database, an online tool for recording, analyzing and disseminating virus-host interactions., Bioinformatics 2021 Feb; (): .
-
2021biotoolsSchema: a formalized schema for bioinformatics software description., Gigascience 2021 Jan; 10(1): .
-
2021The iPPI-DB initiative: a community-centered database of protein-protein interaction modulators., Bioinformatics 2021 Apr; 37(1): 89-96.
-
2021Perspectives on automated composition of workflows in the life sciences., F1000Res 2021 ; 10(): 897.
-
2021Bioimage analysis workflows: community resources to navigate through a complex ecosystem., F1000Res 2021 ; 10(): 320.
-
+View full list of publications