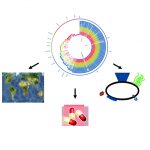
Previously controlled bacterial infections can re-emerge due to antibiotic resistance or vaccine escape. Species of pathogenic bacteria comprise a huge amount of genotypic and phenotypic diversity. Our lab is interested in the diversity, evolution and epidemiology of bacterial pathogens and in the links between the genotypic and phenotypic (ecology, colonization, transmission, virulence, antibiotic resistance, immune response) diversity of the strains within particular species. We focus on three pathogens of high public health importance: the multidrug resistant Klebsiella pneumoniae, which causes various types of infections including urinary tract, respiratory and blood infections; Bordetella pertussis, the agent of whooping cough; and Corynebacterium diphtheriae, the agent of diphtheria. We combine epidemiological surveillance, microbiology, genomics, proteomics, bioinformatics and immunological approaches as well as in-vivo and in-vitro models of infection. We also develop and maintain genomic libraries of bacterial genotypes and strain nomenclatures that facilitate global collaborative surveillance and population biology of bacterial pathogens.
Click to view graph
Connections
About
Members
Former Members
2000
2000
Name
Position
2017
2019
Marina Barros
Post-doc
2017
2019
Melody Dazas
Technician
2017
2019
Sandra Corre
Research Engineer
2017
2019
Leonardo-Gabriel Panunzi
Post-doc
2017
2019
Juan Sebastián López Fernández
Post-doc
2017
2019
Nadine Pouradier
Technician
2017
2019
Pierre Tonnerre
Post-doc
2017
2019
Pauline Leroux
Graduate Student
2016
2022
Sophie Guillot
Deputy-Head NRC Whooping Cough
2020
2023
Sebastien Bridel
Post-doc
2016
2022
Isabelle Moulherat
Assistant
2020
2023
Soraya Matczack
PhD student
2018
2023
Melanie Hennart
PhD Student and post-doc
2016
2023
Annick Carmi-Leroy
Technician