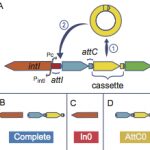
The UMR3525 studies several aspects of genome biology, with a focus on maintenance and evolution of genetic information. On one hand, most teams work on the organisation, expression, and replication of genomes. On the other hand, most teams study how genomes change through time by mutations, gene acquisition, DNA loss and genetic rearrangements. The key biological models are enterobacteria, Vibrio, and yeast. Recently, the developments in our labs and the integration of the Berthelot team spurred development of projects in humans (and other mammals).
Click to view graph
Connections
Members
Groups
Laboratory
Dynamics of the Genome

Benoit Arcangioli
Laboratory
Bacterial Genome Plasticity

Didier Mazel
Laboratory
Microbial Evolutionary Genomics

Eduardo Rocha
Laboratory

Spatial Regulation of Genomes

Romain Koszul
Laboratory
UMR
G5

Comparative Functional Genomics

Camille Berthelot
Group

Group: Guy-Franck Richard

Guy-Franck Richard
Laboratory
G5

Molecular diversity of microbes

Aude Bernheim
Former Members
Former directors of the UMR3525 or URA2171:
- Alain Jacquier (2012-2022)
- Bernard Dujon (2000-2011)
Former principal investigators of the UMR3525 or URA2171:
- Alain Jacquier (Genetics of molecular interactions)
- Bernard Dujon (Molecular genetics of yeast, retired)
- Antoine Danchin (Genetics of bacterial genomes, retired)
- Massimo Vergassola (Physics of the biological systems)
- Frank Kunst (Bacterial genomics, deceased)
- Carmen Buchrieser (Biology of intracellular bacteria)
- Philippe Glaser (Ecology and Evolution of Antibiotics Resistance)
- Roland Brosch (Integrated Mycobacterial Pathogenomics)
Software
Fundings
Publications
Download-
2024Type IV-A3 CRISPR-Cas systems drive inter-plasmid conflicts by acquiring spacers in trans., Cell Host Microbe 2024 May; (): .
-
2024Sister chromatid cohesion halts DNA loop expansion., Mol Cell 2024 Mar; 84(6): 1139-1148.e5.
-
2024Perturbation and resilience of the gut microbiome up to 3 months after β-lactams exposure in healthy volunteers suggest an important role of microbial β-lactamases., Microbiome 2024 Mar; 12(1): 50.
-
2024Capsules and their traits shape phage susceptibility and plasmid conjugation efficiency, Nature Communications, 2024, 15 (1), pp.2032. ⟨10.1038/s41467-024-46147-5⟩.
-
2024Phage-plasmids promote recombination and emergence of phages and plasmids., Nat Commun 2024 Feb; 15(1): 1545.
-
2024Orchestrating chromosome conformation capture analysis with Bioconductor., Nat Commun 2024 Feb; 15(1): 1072.
-
2024Cassette recombination dynamics within chromosomal integrons are regulated by toxin-antitoxin systems., Science Advances , 2024, 10 (2), pp.eadj3498. ⟨10.1126/sciadv.adj3498⟩.
-
2024Transcription-induced domains form the elementary constraining building blocks of bacterial chromosomes., Nat Struct Mol Biol 2024 Jan; (): .
-
2024Integron cassettes integrate into bacterial genomes via widespread non-classical attG sites., Nat Microbiol 2024 Jan; 9(1): 228-240.
-
2023Chance Favors the Prepared Genomes: Horizontal Transfer Shapes the Emergence of Antibiotic Resistance Mutations in Core Genes., Mol Biol Evol 2023 Oct; 40(10): .
-
+View full list of publications