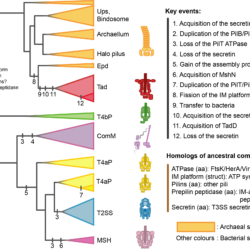
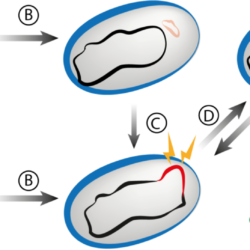
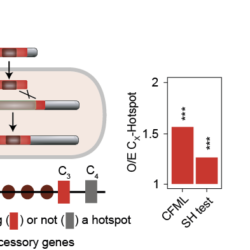
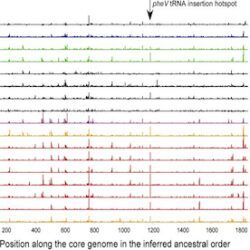
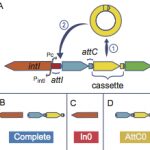
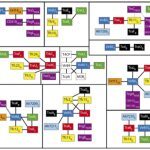
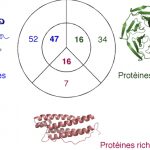
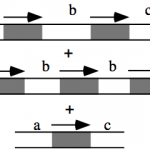
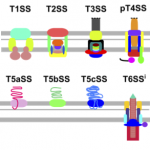
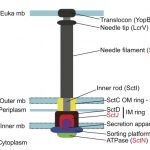
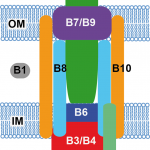
My work focuses on the interplay between the organisation of genomes arising by the chromosomal interactions of cellular processes and the recombination processes generating adaptive genetic variation. Genetic elements tend to be organized in genomes by natural selection in order to optimize this highly complex structure. Yet, they are also constantly faced with genomic variations including mutations, amplifications, rearrangements and horizontal gene transfer. I study the conflict between organisational features and the adaptive plasticity of genomes and how it determines genome evolution and ecological adaptation.
About
Transversal Projects
Projects
Software
Tools
CV
Since 2023 – Head of the UMR3525 CNRS/Institut Pasteur
2019-2023 – Head of the Genomes & Genetics department at Pasteur Institute.
Since 2013 – Head of the unit Microbial Evolutionary Genomics at Pasteur Institute.
Since 2009 – Senior researcher (DR2 then DR1) at the UMR3525 CNRS in Pasteur Institute.
2016-2017 – Member of the COMESP (president SC5)
2015-17 – Associate director for research of the C3BI (Pasteur Institute).
2012-16– President of the CID51 of the National Research Committee (CoNRS).
2010-2015 – Associate director of the GdR BIM.
2008 to 2012– Head junior team Microbial Evolutionary Genomics in URA2171 and UMR3525.
2005– Habilitation in Biology (U Pierre et Marie Curie, Paris).
2000 to 2009– Junior researcher (CR) at the URA2171 CNRS in the Atelier de BioInformatique at UPMC and at Pasteur Institute.
2000– PhD in BioInformatics (U Versailles Saint–Quentin en Yvelines, France).
1997– MSc in Applied Mathematics (Lisbon Technical U, Portugal).
1995– BSc and MSc in Chemical Engineering/Biotechnology (Lisbon Technical U, Portugal).
Publications
Download-
2024Type IV-A3 CRISPR-Cas systems drive inter-plasmid conflicts by acquiring spacers in trans., Cell Host Microbe 2024 May; (): .
-
2024Perturbation and resilience of the gut microbiome up to 3 months after β-lactams exposure in healthy volunteers suggest an important role of microbial β-lactamases., Microbiome 2024 Mar; 12(1): 50.
-
2024Capsules and their traits shape phage susceptibility and plasmid conjugation efficiency, Nature Communications, 2024, 15 (1), pp.2032. ⟨10.1038/s41467-024-46147-5⟩.
-
2024Phage-plasmids promote recombination and emergence of phages and plasmids., Nat Commun 2024 Feb; 15(1): 1545.
-
2024Cassette recombination dynamics within chromosomal integrons are regulated by toxin-antitoxin systems., Science Advances , 2024, 10 (2), pp.eadj3498. ⟨10.1126/sciadv.adj3498⟩.
-
2024Integron cassettes integrate into bacterial genomes via widespread non-classical attG sites., Nat Microbiol 2024 Jan; 9(1): 228-240.
-
2023Chance Favors the Prepared Genomes: Horizontal Transfer Shapes the Emergence of Antibiotic Resistance Mutations in Core Genes., Mol Biol Evol 2023 Oct; 40(10): .
-
2023Latent evolution of biofilm formation depends on life-history and genetic background, npj Biofilms and Microbiomes, 2023, 9 (1), pp.53. ⟨10.1038/s41522-023-00422-3⟩.
-
2023Competition between lysogenic and sensitive bacteria is determined by the fitness costs of the different emerging phage-resistance strategies., Elife 2023 Mar; 12(): .
-
2023Identification and characterization of thousands of bacteriophage satellites across bacteria., Nucleic Acids Res 2023 Mar; (): .
-
+View full list of publications