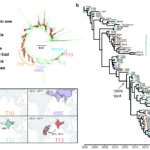
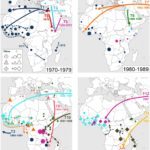
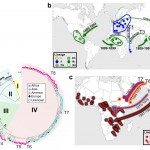
The Unit includes the National Reference Center (CNR) for Salmonella, Shigella/Escherichia coli and the NRC for Vibrio and Cholera, and a WHO Collaborating Center. Our research interests are strongly linked to public health and reference activities for the enteric bacterial pathogens monitored by the Unit. We work on three partly overlapping themes: (i) identification and dynamics of bacterial populations resistant to antibiotics (ii) development of new bacterial typing and diagnostic tools, (iii) population structures and the evolution of genetically monomorphic pathogenic agents, with a special focus on emergent, epidemic and antimicrobial-resistant bacterial populations.
Click to view graph
Connections
Members
Former Members
Courses
Waterborne Infectious Diseases MOOC
This MOOC explains why water can transmit bacterial, viral and parasitic infections and explores means of control and prevention. Register on: https://www.fun-mooc.fr/en/cours/water-borne-infectious-diseases/ Enrollment: From Oct.5, 2023 to Feb. 7, 2024 Course: From Dec.5, 2023 to Feb. […]
2023-12-05 00:00:00
2024-02-14 23:00:00
Europe/Paris
Waterborne Infectious Diseases MOOC
This MOOC explains why water can transmit bacterial, viral and parasitic infections and explores means of control and prevention. Register on: https://www.fun-mooc.fr/en/cours/water-borne-infectious-diseases/ Enrollment: From Oct.5, 2023 to Feb. 7, 2024 Course: From Dec.5, 2023 to Feb. […]
Projects
Transversal Project
Tools
Fundings
Partners
Publications
Download-
2023The evolution and international spread of extensively drug resistant Shigella sonnei, Nat Commun 14, 1983 (2023).
-
2023Editorial: Mobile genetic elements as dissemination drivers of multidrug-resistant Gram-negative bacteria., Front Cell Infect Microbiol 2023 ; 13(): 1180510.
-
2023Rapid emergence of extensively drug-resistant Shigella sonnei in France., Nat Commun 2023 Jan; 14(1): 462.
-
2023Countrywide multi-serotype outbreak of Salmonella Bovismorbificans ST142 and monophasic Salmonella Typhimurium ST34 associated with dried pork sausages in France, September 2020* to January 2021., Euro Surveill 2023 Jan; 28(2): .
-
2022Excess risk of subsequent infection in hospitalized children from a community cohort study in Cambodia and Madagascar., Int J Epidemiol 2022 Oct; 51(5): 1421-1431.
-
2022Clinical and experimental bacteriophage studies: Recommendations for possible approaches for standing against SARS-CoV-2., Microb Pathog 2022 Mar; 164(): 105442.
-
2022Population structure analysis and laboratory monitoring of Shigella by core-genome multilocus sequence typing., Nat Commun 2022 Jan; 13(1): 551.
-
2022Leveraging serology to titrate immunisation programme functionality for diphtheria in Madagascar., Epidemiol Infect 2022 Jan; 150(): e39.
-
2021Azithromycin resistance in Shiga-toxin Producing Escherichia coli in France between 2004 and 2020 and detection of mef(C)-mph(G) genes., Antimicrob Agents Chemother 2021 Dec; (): AAC0194921.
-
2021Improved molecular diagnosis and culture of the emerging heteropathotype enterohemorrhagic Escherichia coli O80:H2 using its non-melibiose-fermenting and antibiotic-resistance properties., J Clin Microbiol 2021 Sep; (): JCM0153021.
-
+View full list of publications
Contact
Enteric Bacterial Pathogens Unit 28 rue du Docteur Roux -75724 PARIS Cedex 15-France. Tel : + 33 (1) 45 68 83 45 (secretary: + 33 (1) 83 92 21 98) Fax : + 33 (1) 45 68 88 37 E-mail : francois-xavier.weill@pasteur.fr