The Computational Systems Biomedicine lab builds computational approaches to discover causal paths in complex diseases that connect their genetic or environmental causes to their higher-level effects.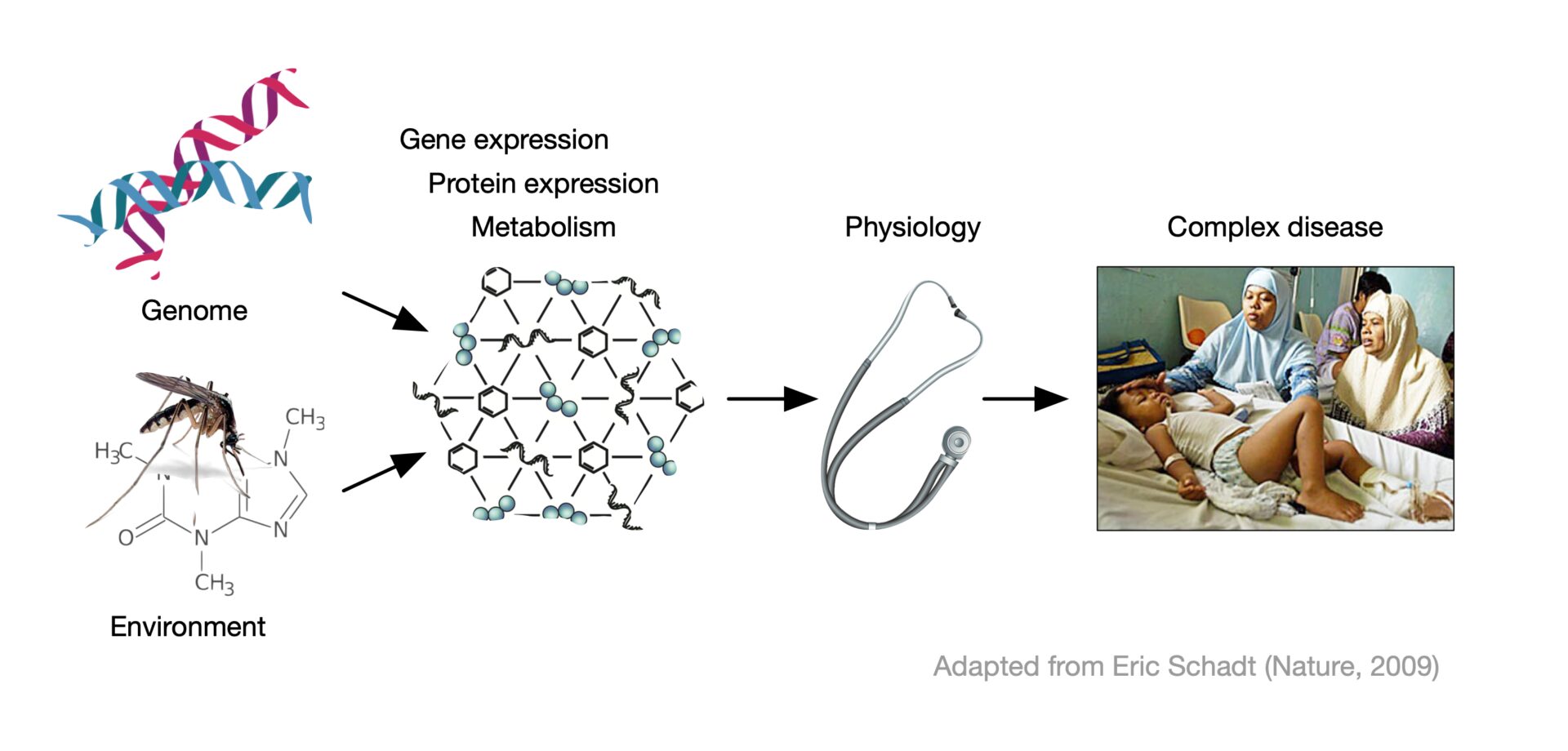
Previous research
The rapid development of new measurement technologies provides unprecendented opportunities to observe complex disease at a large scale, and at different levels, including molecular (e.g., transcriptomic, proteomic, and metabolomic) networks. Much of our research is therefore based on machine learning tools. Better understanding of the causal disease paths will lead to better treatment choices, and, ultimately, to medicine that prevents disease instead of reacting to it. Our group is probably best known for discovering the ‘guilt-by-association’ principle in molecular networks, and for creating Cytoscape, an open-source community platform to visualize and analyze molecular networks. We are also member of the national “laboratory of excellence” (LabEx) programs Milieu Intérieur and IBEID.
Future projects and goals
New developments in this very dynamic area of research are rapidly evolving from three interrelated aspects:
- new measurement technologies (such as single cell RNA sequencing; Blecher-Gonen et al. 2019), requiring
- new computational tools (such as Viral-Track; Bost et al. 2020 and LEAN analysis; Gwinner et al. 2017), leading to
- new biological insights (such as the detailed characterization of COVID 19-infected cells Bost et al. 2020, and a sparse, reproducible, biomarker for severe dengue infection; Nikolayeva et al. 2018), which then leads to new biological questions that need to be addressed with their own measurement and computational tools.
We therefore couple the development of new computational tools closely to the development of new technology, and ambitious biomedical research projects, currently in the areas of single cell sequencing technology, autoimmune disease and cancer. If you might be interested to join our research team, please check out the Jobs section below or contact us.