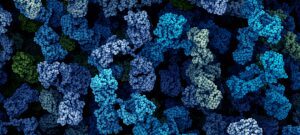
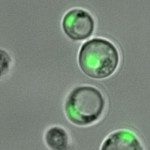
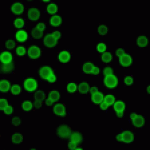
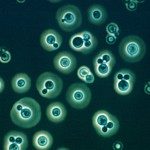
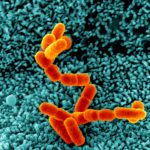
The project of the Unit RNA Biology of Fungal Pathogens focuses on the study of the structure and plasticity of the transcriptomes of human fungal pathogens. We primarily utilize the basidiomycete yeast Cryptococcus neoformans as a model, but we also consider additional pathogenic or non-pathogenic fungi for comparison. We employ diverse OMICs strategies to investigate the diversity of mRNA and lncRNA isoforms present in the fungal cell. Additionally, we are interested in understanding the mechanisms that regulate this diversity and its impact on the biology of fungal cells. These foundational findings are applied to more virulence-oriented questions, exploring how the transcriptome structure is regulated during infection and the consequences of these modifications for both the fungus and the host.
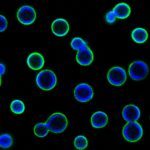
More recently, we have initiated research into the extracellular vesicles (EVs) produced by fungi. Our project delves into the structure and dynamics of the vesicular transcriptome, as well as the study of EV biosynthetic pathways. We are also investigating the roles of these extracellular organelles in cell-to-cell communication and host-pathogen interactions. This second part of the project is also a component of the Pasteur international Unit “Fungal Extracellular Vesicles,” led in collaboration with Robin May (University of Birmingham, UK) and Marcio Rodrigues (Fiocruz, Brazil).