Development and harmonization of innovative methods for comprehensive analysis of food-borne toxigenic bacteria, i.e. Staphylococci, Bacillus cereus and Clostridium perfringens.
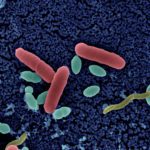
The bacterial toxins produced by the strains of Staphylococcus, Clostridium and Bacillus are responsible for food poisoning (about 10,000 cases per year in the European Union with certainly an underestimation of cases for many reasons). In this context, the project aims to develop non-NGS approaches to detect and quantify the toxins involved (LC MS / MS, MALDI-ToF and ELISA), propose methods to differentiate pathogenic strains from non-pathogenic strains, and perform a series of inter-laboratory tests to evaluate and optimize the performance of the methods developed. The CIP operates at different levels of the project, S. aureus and B. cereus in particular, with Dominique Clermont as responsible for a work package.
Antibiotic resistance: emergence and gene transfer
The emergence of antibiotic resistance creates a new challenge for public health, and there is no simple solution. CIP is hosting a unique collection of clinical isolates to study history and evolution of resistance in ESKAPE pathogens (Enterococcus faecium, Staphylococcus aureus, Klebsiella pneumoniae, Acinetobacter baumanii, Pseudomonas aeruginosa) and many others.
Resistance to antibiotics can be either intrinsic to a given bacterial species or acquired by genetic mutations and foreign DNA pieces not present in the natural populations. Species identification and means to identify genes coding for resistance to antibiotics are integrated in our pipelines of genome data analysis from CIP deposited strains. We are developing sophisticated methods to implement such a resistance gene prediction workflow in order to detect what is not already known, searching all possible acquired resistance genes and novel resistance signatures.
Genome sequencing of type strains
The CIP, because of the great biodiversity of its collection, was requested by the WDCM (WFCC-MIRCEN World Data Center for Microorganisms) (http://www.wdcm.org/) to participate in a large-scale genome sequencing project of type strains over five years.
Valorization of the biotechnological potential of CIP strains
200 strains of bacteria belonging to the group Bacillus cereus sensu lato were studied in terms of belonging to this or that species of the group, their content of toxins or virulence factors. The analysis of the genomes also made it possible to detect clusters of interest. The characterization of unknown molecules with potential antimicrobial activity is ongoing.