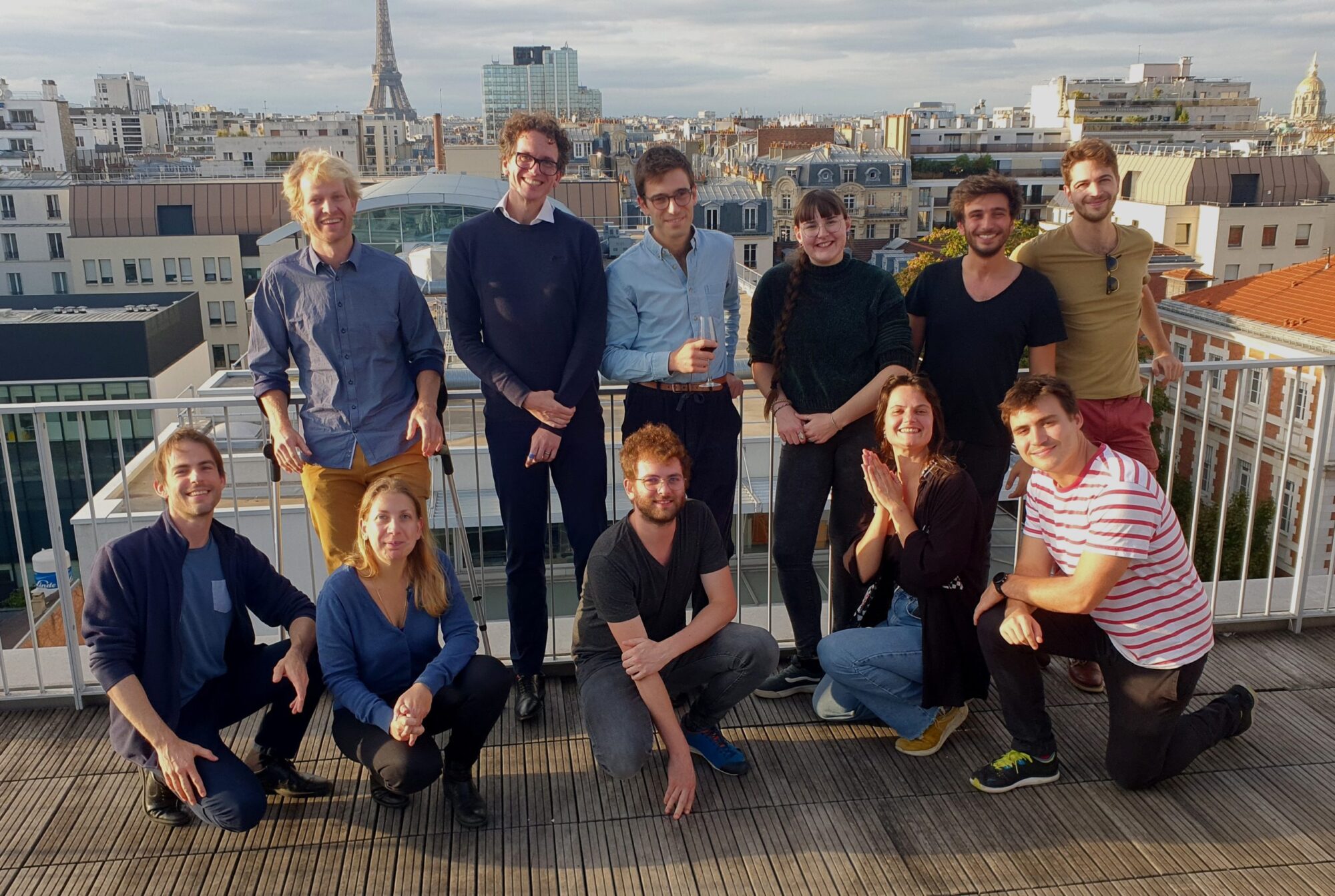
The lab is focused on the algorithms and computation selected by evolution to perform biological decision-making. We address this topic with an interdisciplinary approach mixing statistical physics, Bayesian machine learning, information theory and various experimental biological setups.
If you have inquiries about open positions, please write to dbc-epi-recrutement at pasteur dot fr