About
This project aims to characterize the mircrociruit structure of small animals’ brains and link this to how the brain processes information and encodes behavior. The project relies on simultaneous access to recent synapse-resolution wiring diagrams of complete brain regions and single-neuron optogenetic control in freely behaving animals. We employ methods from graph theory, statistical learning, and information theory to characterize the topology and structure of microcircuits in animal brains and to link the circuits’ structures to how they perform decision making and generate behavioral sequences.
A prominent hypothesis for how the brain’s wiring is organized is that it is made up of stereotyped “canonical” microcircuits. Relying on a restricted set of repeated circuits, optimized for specific functions, may provide biologically advantageous inductive biases for efficient learning and help encode innate behaviors. Information theory furthermore tells us that the presence of statistically regular circuits makes the connectome compressible. Canonical microcircuits thus provide a means that evolution may have selected to encode the neural wiring information more efficiently in the limited storage space of the genome.
Extracting the statistical regularities (“motifs”) of a connectome is a challenging inverse problem since one often has access to only a single experimental realization (i.e., a single graph). The analysis traditionally relies on a statistical model against which statistical significance is defined. The choice of this null model is ambiguous however, leading to possible uncontrolled biases. Furthermore, the number of possible microcircuit motifs increases combinatorially with their size, making motif mining a multiple testing problem.
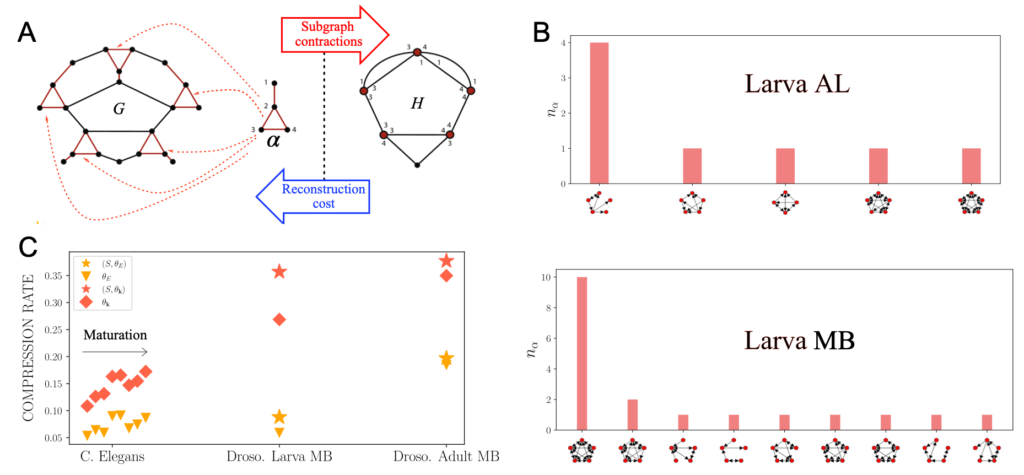
Relying on the equivalence between compression and statistical learning, we develop lossless network compression techniques based on subgraph contractions and subgraph covers. These allow us to find combinations of motifs that maximally compress a connectome and evaluate their combined significance, circumventing problems related to multiple testing encountered when mining motifs individually using null hypothesis testing, and compare the compressibility of different connectomes.