Link to Pubmed [PMID] – 23675309
PLoS Genet. 2013 May;9(5):e1003493
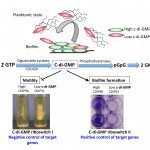
Clostridium difficile is an emergent pathogen, and the most common cause of nosocomial diarrhea. In an effort to understand the role of small noncoding RNAs (sRNAs) in C. difficile physiology and pathogenesis, we used an in silico approach to identify 511 sRNA candidates in both intergenic and coding regions. In parallel, RNA-seq and differential 5′-end RNA-seq were used for global identification of C. difficile sRNAs and their transcriptional start sites at three different growth conditions (exponential growth phase, stationary phase, and starvation). This global experimental approach identified 251 putative regulatory sRNAs including 94 potential trans riboregulators located in intergenic regions, 91 cis-antisense RNAs, and 66 riboswitches. Expression of 35 sRNAs was confirmed by gene-specific experimental approaches. Some sRNAs, including an antisense RNA that may be involved in control of C. difficile autolytic activity, showed growth phase-dependent expression profiles. Expression of each of 16 predicted c-di-GMP-responsive riboswitches was observed, and experimental evidence for their regulatory role in coordinated control of motility and biofilm formation was obtained. Finally, we detected abundant sRNAs encoded by multiple C. difficile CRISPR loci. These RNAs may be important for C. difficile survival in bacteriophage-rich gut communities. Altogether, this first experimental genome-wide identification of C. difficile sRNAs provides a firm basis for future RNome characterization and identification of molecular mechanisms of sRNA-based regulation of gene expression in this emergent enteropathogen.