Directeur de Recherche, CNRS
Click to view graph
Connections
Click to view timeline
Timeline
About
Projects
Tools
Patents
Former Teams
Publications
Download-
2024Structure and Dynamics of Type 4a Pili and Type 2 Secretion System Endopili, In: Macromolecular Protein Complexes V, vol 104. Chapter 21. p549-563.
-
2024Mechanism of action of phthalazinone derivatives against rabies virus., Antiviral Res 2024 Apr; 224(): 105838.
-
2024Bonds and bytes: The odyssey of structural biology, Curr Opin Struct Biol. 2024 Feb;84:102746. doi: 10.1016/j.sbi.2023.102746. .
-
2024Ton motor conformational switch and peptidoglycan role in bacterial nutrient uptake., Nat Commun 2024 Jan; 15(1): 331.
-
2023The Mla system of diderm Firmicute Veillonella parvula reveals an ancestral transenvelope bridge for phospholipid trafficking., Nat Commun 2023 Nov; 14(1): 7642.
-
2023A poly-proline II helix in YadA from Yersinia enterocolitica serotype O:9 facilitates heparin binding through electrostatic interactions., FEBS J 2023 Nov; (): .
-
2023Structural Analysis of Proteins from Bacterial Secretion Systems and Their Assemblies by NMR Spectroscopy, Bacterial Secretion Systems, 2715, Springer US, pp.503-517, 2023, Methods in Molecular Biology, ⟨10.1007/978-1-0716-3445-5_30⟩.
-
2023Structure and dynamic association of an assembly platform subcomplex of the bacterial type II secretion system., Structure 2023 Feb; 31(2): 152-165.e7.
-
2022Secondary structure and 1H, 15 N & 13C resonance assignments of the periplasmic domain of OutG, major pseudopilin from Dickeya dadantii type II secretion system., Biomol NMR Assign 2022 Apr; (): .
-
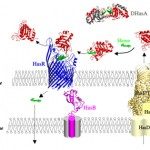
2022Structural and molecular determinants for the interaction of ExbB from Serratia marcescens and HasB, a TonB paralog, Commun Biol . 2022 Apr 13;5(1):355. .
-
+View full list of publications