CR CNRS
Click to view graph
Connections
Click to view timeline
Timeline
Courses
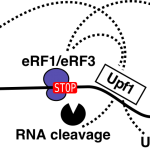
Multiple roles of RNAs
A practical course on RNA methods RNA research is at the basis of our current understanding of gene expression regulation and was crucial for the discovery of tools such as siRNA and CRISPR/Cas9. RNA […]
2023-02-13 09:00:00
2023-02-24 18:00:00
Europe/Paris
Multiple roles of RNAs
A practical course on RNA methods RNA research is at the basis of our current understanding of gene expression regulation and was crucial for the discovery of tools such as siRNA and CRISPR/Cas9. RNA […]
28 Rue du Docteur Roux, Paris, France
Projects
Former Teams
Publications
Download-
2023Deadenylation rate is not a major determinant of RNA degradation in yeast, bioRxiv, 2023.01.16.524186.
-
2018Antisense transcriptional interference mediates condition-specific gene repression in budding yeast, Nucleic Acids Res. 2018 May;.
-
2015Systematic Determination of Transcription Factor DNA-Binding Specificities in Yeast, Methods Molecular Biology, Vol. 1361, Frédéric Devaux (Eds): Yeast Functional Genomics, 978-1-4939-3078-4, 318712_1_En, (12).
-
2013Yeast ribosomal protein L7 and its homologue Rlp7 are simultaneously present at distinct sites on pre-60S ribosomal particles, Nucleic Acids Res. 2013 Nov;41(20):9461-70.
-
2010Objective sequence-based subfamily classifications of mouse homeodomains reflect their in vitro DNA-binding preferences, 2010 Dec;38(22):7927-42..
-
2010Genome-wide analysis of ETS-family DNA-binding in vitro and in vivo, EMBO J. 2010 Jul 7;29(13):2147-60.
-
2009Diversity and complexity in DNA recognition by transcription factors, Science. 2009 Jun 26;324(5935):1720-3..
-
2009TFCat: the curated catalog of mouse and human transcription factors, Genome Biol. 2009;10(3):R29.
-
2008A library of yeast transcription factor motifs reveals a widespread function for Rsc3 in targeting nucleosome exclusion at promoters, Mol Cell. 2008 Dec 26;32(6):878-87..
-
2008Predicting the binding preference of transcription factors to individual DNA k-mers, Bioinformatics 2009 Apr;25(8):1012-8.
-
+View full list of publications