Our research interests are mainly focused on the structure-function relationships of proteins (in particular crucial enzymes belonging to the nucleotide metabolism) through multidisciplinary approaches involving mainly biochemistry but also structural biology and genetics. Our studies have been concentrated on nucleoside monophosphate kinases (NMPKs) and more recently IMP dehydrogenase (IMPDH): see below for more details in the project section.
Related to the validation of these proteins as potent targets for the development of new anti-infectious agents, another aspect of our activity concerns the screening of chemical compounds. The screening activity (acronym CBC for “Criblage Biologique et Chimiothèques”) is devoted to investigating the effect of small molecules mainly in the field of anti-infectious agents, in our research programs or through collaborative projects.
The offered resources are assistance in assay development and validation, as well as actual screening and data processing. Our compound collection comprises commercial and academic chemical libraries (around 71,000 compounds in July 2015). One third of the collection is composed of commercial chemical libraries acquired through Prestwick Chemical (1,200 compounds; www.prestwickchemical.com), ChemDiv (9,360 compounds from a kinase inhibitor subset library and 4,624 from a nucleobase analog subset library; eu.chemdiv.com) and Chem-X-Infinity (10,000 compounds as a representative subset of their collections) companies. The rest of the collection is from the French “Chimiothèque Nationale” (http://www.chimiotheque-nationale.org/) including the chemical libraries from the Chemistry and Biocatalysis Unit of the Institut Pasteur, the Institut Curie, the Institut de Chimie et Biochimie Moléculaires et Supramoléculaires (ICBMS; Lyon), the Institut de Chimie des Substances Naturelles (ICSN; Gif-sur-Yvette), the Faculté de Pharmacie de Strasbourg, the Centre d’Etude et de Recherche sur le Médicament de Normandie (CERMN; Caen), the Institut de Chimie Organique et Analytique (ICOA; Orléans), BioCIS (Faculté de Pharmacie, Chatenay-Malabry) and LCBPT (Université Paris Descartes). Since the set up, 20 campaigns have been performed in our research programs (UMPKmt, IMPDH) or by means of collaborative projects.
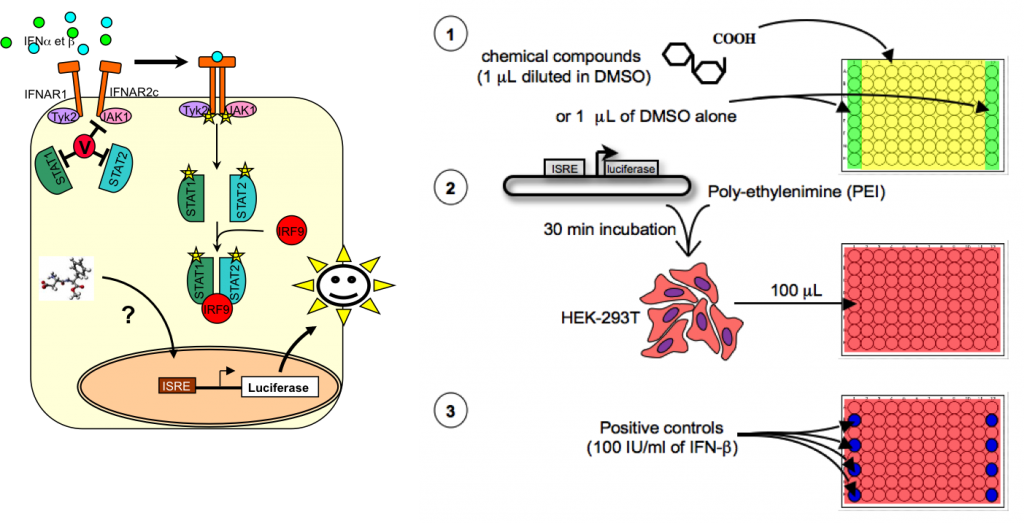
Our first collaborative project, namely “STING” for “Stimulators of Interferon-inducible Genes”, was developed with the Viral Genomics and Vaccination Unit (coordinator P.O. Vidalain; funded by the Institut Carnot Pasteur MI). The goal of this project was to identify molecules stimulating the innate antiviral response, in particular the expression of IFN-Stimulated Genes (ISGs; Vidalain et al, 2015). A cellular assay was developed to screen chemical libraries, and select activators of ISGs (Lucas-Hourani et al, 2014).
This assay was based on human cells transfected with a luciferase reporter gene under control of type I interferon-stimulated response elements (ISRE), and was used in a high-throughput setting to screen a chemical library comprising 54,000 compounds (see figure above). Among validated hits, compound DD264 displayed a broad-spectrum antiviral activity. While deciphering its mode of action, we demonstrated that DD264 blocks de novo pyrimidine biosynthesis by inhibiting the fourth enzyme of this pathway, namely dihydroorotate dehydrogenase (DHODH). Altogether, these results established for the first time a functional link between innate immunity and cellular levels of pyrimidine nucleosides (Lucas-Hourani et al, 2013). We have further established that the antiviral activity of DD264 and brequinar, a well-characterized DHODH inhibitor, was strictly dependent on transcription of cellular genes, export of mRNA from nucleus and on IRF1 transcription factor. Thus, the antiviral activity of pyrimidine biosynthesis inhibitors is not a direct consequence of the viral machinery running dry of pyrimidines but rather relies on stimulation of the innate immune response. Another chemical family (Lucas-Hourani et al, 2015; Munier-Lehmann et al, 2015) with the same mode of action, but showing a much higher potency, was more recently developed. Unfortunately, several attempts to use pyrimidine synthesis inhibitors in the treatment of systemic viral infections have failed, probably because high uridine concentrations in peripheral blood (about 3-8 µM) compensate for the inhibition of de novo pyrimidine biosynthesis. We are now looking for other therapeutic applications for these compounds (autoimmune diseases such as rheumatoid arthritis and multiple sclerosis). More recently, a new library was screened using the same assay, and a novel immuno-stimulatory compound, showing a different mode of action, was identified. Its mode of action is currently under investigation.
The main goal of the FUNGI program (coordinator J.P. Latgé; funded by the Institut Carnot Pasteur MI) was the identification of novel lead compounds for the development of new drug combinations to treat systemic fungal infections (collaboration with the Fungal Biology and Pathogenicity Unit, C. d’Enfert and the Aspergillus Unit, J.-P. Latgé). The current drugs against the two main fungal pathogens Candida albicans and Aspergillus fumigatus are insufficiently potent and the combination of drugs is a way to improve their efficacy. Using a resazurin-based assay (Clavaud et al, 2012), we have identified new compounds active against both fungi in combination with known antifungals that were confirmed in secondary assays by checkerboard assays (Zeidler et al, 2013). In the course of these screenings, a novel chemical series active alone on both fungi was identified with low MICs (below 5μg/ml): antifungal activity was never reported for this family. Moreover, the most potent compound of this family did not show any mutagenic activity in vitro, nor toxicity in mice. We are currently investigating the cellular target of this compound family.
We have been part of the FP7 Leisdrug consortium (coordinator G. Spaeth), in particular in WP1. The goal was to identify compounds that selectively kill intracellular Leishmania amastigotes in primary macrophages using a phenotypic screening assay (Aulner et al., 2013).
We have other on-going collaborative projects: i) with the Mycobacterial Genetics Unit (B. Gicquel) with the aim of identifying molecules with antimycobacterial activity. Three different fast growing mycobacterial species easy to manipulate will be used with the hypothesis that escape mechanisms will have not been developed in all of them. ii) with the Viral Pathogenesis Unit (F. Arenzana-Seisdedos) for the identification of CXCR4 or CXCR7 antagonists for interfering in CXCL12-mediated pathologic cell signalling or HIV entry. An assay based on the Tag-lite® technology developed by Cisbio was used to screen the academic chemical libraries. Hits from the CERMN were validated in secondary assays, and a collaborative research program is underway to develop this chemical series.