Cliquez pour voir le graph
Connexions
Cliquez pour voir la ligne de temps
Ligne de temps
Cours
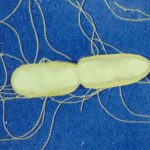
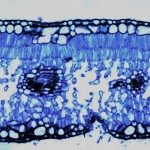
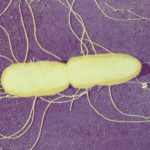
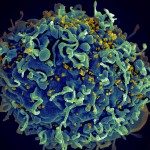
Waterborne Infectious Diseases MOOC
This MOOC explains why water can transmit bacterial, viral and parasitic infections and explores means of control and prevention. Register on: https://www.fun-mooc.fr/en/cours/water-borne-infectious-diseases/ Enrollment: From Oct.2, 2025 to Dec. 1, 2026 Course: From Dec.2, 2025 to Dec. […]
2025-12-02 00:00:00
2026-12-01 23:00:00
Europe/Paris
Waterborne Infectious Diseases MOOC
This MOOC explains why water can transmit bacterial, viral and parasitic infections and explores means of control and prevention. Register on: https://www.fun-mooc.fr/en/cours/water-borne-infectious-diseases/ Enrollment: From Oct.2, 2025 to Dec. 1, 2026 Course: From Dec.2, 2025 to Dec. […]
Projets Transversaux
Projets
Outils
Publications
Télécharger-
2025Modeling the spatio-temporal spread of cholera in France in 1892., Epidemics 2025 Dec; 53(): 100872.
-
2025Genomic characterisation of multidrug-resistant Salmonella enterica serovar Kentucky ST198 isolates from various sources in Algeria, North Africa., Microb Genom 2025 Nov; 11(11): .
-
2025Genomics reveals multiple introductions of the seventh pandemic Vibrio cholerae O1 El Tor lineage into Iran since 1965., Microb Genom 2025 Oct; 11(10): .
-
2025Global diversity and evolution of Salmonella enterica serovar Panama: a genomic epidemiology study., Lancet Microbe 2025 Sep; 6(9): 101150.
-
2025Recent and convergent reversion to serotype Ogawa in the AFR12 sublineage of Vibrio cholerae O1 El Tor in Cameroon., Microb Genom 2025 Sep; 11(9): .
-
2025Genomic census of invasive nontyphoidal Salmonella infections reveals global and local human-to-human transmission., Nat Med 2025 Jul; 31(7): 2325-2334.
-
2025The TyphiNET data visualisation dashboard: unlocking Salmonella Typhi genomics data to support public health., Genome Med 2025 May; 17(1): 51.
-
2025Lessons from 5 Years of Routine Whole-Genome Sequencing for Epidemiologic Surveillance of Shiga Toxin-Producing Escherichia coli, France, 2018-2022., Emerg Infect Dis 2025 May; 31(13): 117-128.
-
2025Salmonella serotypes in the genomic era: simplified Salmonella serotype interpretation from DNA sequence data., Appl Environ Microbiol 2025 Mar; 91(3): e0260024.
-
2024Long-Distance Spread of a Highly Drug-Resistant Epidemic Cholera Strain., N Engl J Med 2024 Dec; 391(23): 2271-2273.
-
+Voir la liste complète de publications