Education
Medical Degree (1996), Cluj, Romania – scientific report on a method for multiple substrate enzyme kinetics parameter estimation. PhD in biochemistry (2001) supervised by Mircea Cucuianu, Cluj, Romania, on the characterisation of proteins through complex purification and mass-spectrometry identification. Habilitation for PhD supervising, HDR (2010) Paris Diderot University.
Honors
Silver medal at the 22nd International Chemistry Olympiad (1990) Thérèse Lebrasseur prize (2019)
Past achievements
(PubMed publications or Google Scholar citations. All my publications or author manuscripts are on HAL-Pasteur)
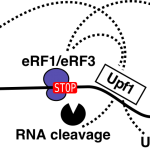
Translation-dependent mRNA degradation in yeast involves unexplored mechanisms and substrates
Genetic interaction profiles are excellent predictors of gene function
In search for new ways to explore gene function, I participated to the development of a method that allows to measure on a large scale the impact on growth of introducing mutants of two different genes in the same strain (Genetic Interactions Mapping, GIM). The best predictor for an unknown gene function can be found by correlating the profiles of sensitivity of mutants to a range of perturbations (PNAS, 2008). A larger scale genome-wide set of screens that I coordinated, led to results that identified new genes involved in RNA transport, maturation and degradation (Nucleic Acids Res, 2021) and triggered several collaborative publications (e.g. Nature Comm 2021, Antimicrob Agents Chemother 2016, Nucleic Acids Res 2014, PNAS 2013).
Ribosome biogenesis occurs through ordered assembly of dozens of proteins
My work on ribosome biogenesis started by with one of the first purification of precursors to the 60S ribosomal subunit, the identification of new factors (EMBO J, 2001) and the first demonstration of an assembly order in this highly complex and conserved pathway (Mol Cell Biol 2003). To establish in which order the 120 factors involved in the formation of 60S ribosomal subunits assemble, I pioneered the use of quantitative mass- spectrometry and combinations of mutants for the study of large complexes assembly (Nucl Acids Res, 2008, 2013).