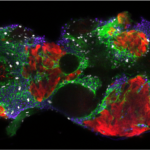
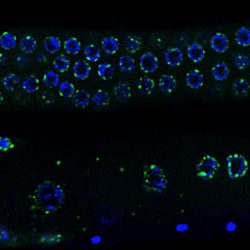
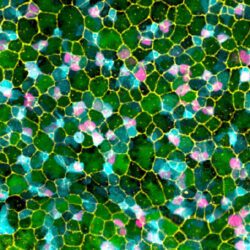
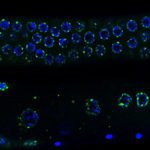
The general objective of our laboratory is to achieve a molecular understanding of the epigenetic inheritance process in animals. 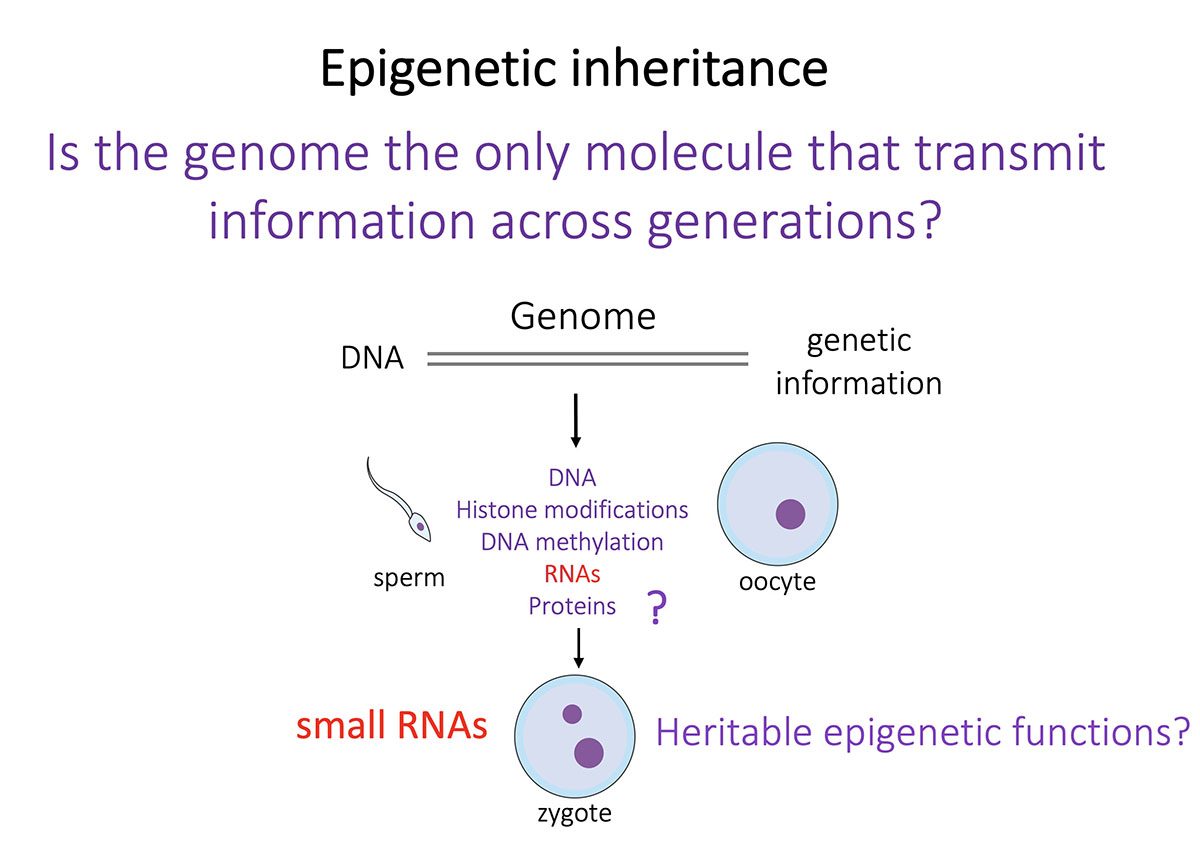 The main goals of our research are i) to elucidate the mechanisms by which small RNAs and histone modifications are used to transmit epigenetic information across generations, ii) to understand how small RNAs are inherited from one generation to another through the gametes, and iii) to explore the mechanisms by which small RNAs can even be transmitted from different tissues and organs to the gametes, to propagate epigenetic information acquired from the soma. To achieve our objective, we are using the nematode Caenorhabditis elegans as an ideal model system to study epigenetic inheritance. WHY C. elegans? The nematode C. elegans is a fertile hermaphrodite, which allows us to obtain a population of worms that are almost genetical identical. Also, its generation time is only three days allowing experiments across multiple generations in only few weeks. Thanks to these features, a number of transgenerational epigenetic phenomena have been documented so far in C. elegans, including the inheritance of traits such as longevity, fertility, and even acquired traits like learning behavior. The existence of this new type of inheritance is certainly a revolutionary idea in biology. However, even if small RNAs and histone modifications have been implicated in these phenomena, the mechanisms underlaying these epigenetic processes still remain very mysterious. We are integrating genetic, biochemical, and molecular biology tools with established high-throughput genomic and proteomic approaches to achieve a deep mechanistic understanding of the role of small RNAs and histone modifications in epigenetic inheritance in animals. We have also optimized biochemical and genomics tools coupled with sophisticated sorting strategies to obtain synchronous populations of individuals, cell types, and organelles at very precise developmental stages. We employ a wide range of cutting edge techniques to study chromatin and epigenetic regulations, which include:
The main goals of our research are i) to elucidate the mechanisms by which small RNAs and histone modifications are used to transmit epigenetic information across generations, ii) to understand how small RNAs are inherited from one generation to another through the gametes, and iii) to explore the mechanisms by which small RNAs can even be transmitted from different tissues and organs to the gametes, to propagate epigenetic information acquired from the soma. To achieve our objective, we are using the nematode Caenorhabditis elegans as an ideal model system to study epigenetic inheritance. WHY C. elegans? The nematode C. elegans is a fertile hermaphrodite, which allows us to obtain a population of worms that are almost genetical identical. Also, its generation time is only three days allowing experiments across multiple generations in only few weeks. Thanks to these features, a number of transgenerational epigenetic phenomena have been documented so far in C. elegans, including the inheritance of traits such as longevity, fertility, and even acquired traits like learning behavior. The existence of this new type of inheritance is certainly a revolutionary idea in biology. However, even if small RNAs and histone modifications have been implicated in these phenomena, the mechanisms underlaying these epigenetic processes still remain very mysterious. We are integrating genetic, biochemical, and molecular biology tools with established high-throughput genomic and proteomic approaches to achieve a deep mechanistic understanding of the role of small RNAs and histone modifications in epigenetic inheritance in animals. We have also optimized biochemical and genomics tools coupled with sophisticated sorting strategies to obtain synchronous populations of individuals, cell types, and organelles at very precise developmental stages. We employ a wide range of cutting edge techniques to study chromatin and epigenetic regulations, which include:
- Global Run On sequencing (GRO-seq)
- Chromatin immunoprecipitation sequencing (ChIP-seq) and Cut and Run
- small RNAs and mRNAs sequencing (RNA-seq)
- Ribosome profiling (Ribo-seq)
- individual-nucleotide resolution Cross-Linking and ImmunoPrecipitation (iCLIP)
The potential capacity of small RNAs to acquire information from different tissues and transmit it across generations would revolutionize biological research and medicine. Therefore, identifying the endogenous RNAs and mechanisms involved in this exchange of information from the soma to the germline and ultimately the next generation could radically change our notion of inheritance. Personal website: www.cecerelab.com