About
Bacteriophages are the most abundant viruses on earth and display an impressive genetic as well as morphologic diversity. Among those, the most common order of phages is the Caudovirales, whose viral particles packages linear double stranded DNA (dsDNA). In this study we investigated how the information gathered by high throughput sequencing technologies can be used to determine the DNA termini and packaging mechanisms of dsDNA phages. The wet-lab procedures traditionally used for this purpose rely on the identification and cloning of restriction fragment which can be delicate and cumbersome. Here, we developed a theoretical and statistical framework to analyze DNA termini and phage packaging mechanisms using next-generation sequencing data. Our methods, implemented in the PhageTerm software, work with sequencing reads in fastq format and the corresponding assembled phage genome.
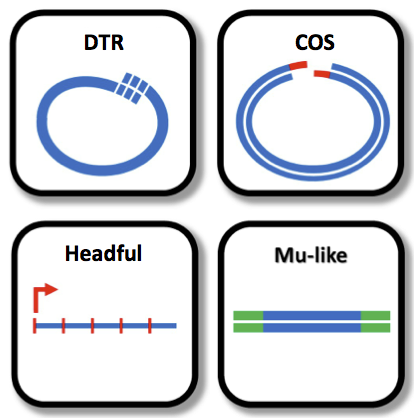
PhageTerm was validated on a set of phages with well-established packaging mechanisms representative of the termini diversity: 5’cos (lambda), 3’cos (HK97), pac (P1), headful without a pac site (T4), DTR (T7) and host fragment (Mu). In addition, we determined the termini of 9 Clostridium difficile phages and 5 phages whose sequences where retrieved from the sequence read archive (SRA).
A direct graphical interface is available as a Galaxy wrapper version at https://galaxy.pasteur.fr and a standalone version is accessible at https://sourceforge.net/projects/phageterm/.