
Scientific computing [133]


Biodiversity and Epidemiology of Bacterial Pathogens
Sylvain Brisse
Numerical methods for network analysis
Christian L. Vestergaard

Vincent Laville
Bioinformatics and Biostatistics HUB
I am a research engineer expert in analyzing large genomic and transcriptomic datasets to decipher the architecture of complex phenotypes. I was recruited by the Hub of Bioinformatics and Biostatistics in 2020 as a […]

Virgile Andreani
Physicist and computer scientist by training, I develop quantitative population models of antibiotic resistance, which I calibrate using optimal experimental design. I then exploit them to engineer optimal treatment strategies.

Christian L. Vestergaard
Decision and Bayesian Computation – Epiméthée
I am a statistical physicist and biophysicist with a main interest in data-driven modeling and in developing methods for the analysis of complex dynamical systems. The focus of my research has been on time-varying […]

InBio: Experimental and Computational Methods for Modeling Cellular Processes
Grégory Batt

François Laurent
Decision and Bayesian Computation – Epiméthée
Biomolecule random walk analysis I am in charge of developping the TRamWAy Python library for random walk analysis. The library features multiple tools to build processing chains/pipelines for the spatial or spatio-temporal resolution of […]

Quentin Giai Gianetto
Bioinformatics and Biostatistics HUB
Since September 2016, I am a research engineer in the Bioinformatics and Biostatistics HUB of the Institut Pasteur and detached in the Proteomics facility. I have a PhD in Signal Processing from the Ecole […]

Thomas Bigot
Expertise group : PAGE – Protein And Gene Evolution
As a bioinformatics engineer, I design analysis methods to extract sequences of known or novel pathogens from metagenomic sequencing data of animal or clinical samples. My work involves evaluating existing methods based on local […]
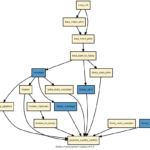
Sequana: a set of flexible NGS pipelines
Thomas Cokelaer

INCEPTION – Institut Convergence for the study of Emergence of Pathology Through Individuals and Populations
IINCEPTION Goal The Inception aims to develop a core structure to mobilize data resources, numerical sciences, and fundamental experimental biology in various health issues (Official website here: https://www.inception-program.fr/en). The inception program uses Integrative Biology, […]

HIV-1 Drug Resistance Mutation analysis
Anna Zhukova

Thomas Obadia
Expertise group : Stats
Thomas is a biostatistician who holds an engineering degree in Agronomy (Agrocampus Ouest, Rennes, France). He also holds a Ph.D. in biostatistics from Université Pierre et Marie Curie for his work on the spread of nosocomial pathogens […]

Olivier Sperandio
Chemoinformatics and proteochemometrics

Anastasia Gazi
Ultrastructural BioImaging
Viruses were for quite long characterized as the invisible heteronomous agents, (unseen by light microscopy & unable to propagate themselves in the absence of susceptible cells) capable of escaping the bacteriological filters (Rivers, 1932, […]