Click to view graph
Connections
Click to view timeline
Timeline
About
Projects
CV
- 2017-to date. DR, Group leader, Biochemistry of Macromolecular Interactions Unit (headed by Daniel Ladant), Institut Pasteur, Paris, France.
- 2014-2016. Head of laboratory of Macromolecular Systems and Signaling, Institut Pasteur, Paris, France.
- 2008-2014. Group leader, Molecular Genetics Unit, dir. by Tony Pugsley, Institut Pasteur, Paris, France. Protein secretion in bacteria.
- 2008. HDR, University Paris XI, Orsay, France.
- 2001-2007. CR, Molecular Genetics Unit, dir. Tony Pugsley, Institut Pasteur, Paris, France.
- 1995-2000. Postdoctoral fellow (Howard Cantarini and FRM), Molecular Genetics Unit, dir. Tony Pugsley. Protein secretion in Escherichia coli.
- 1990-1994. Postdoctoral fellow, Carol Kumamoto laboratory, Department of Microbiology and Molecular Biology, Tufts University Medical School, Boston, USA. Genetic and biochemical analysis of the Escherichia coli Sec system.
- 1987-1990. PhD training in Molecular Biology, Institut for Moelcular Genetics and Genetic Engineering, laboratory of Pr Vladimir Glisin, Belgrade Yugoslavia.
- 1983-1987. Masters of Science, Biochemistry and Molecular Biology, University of Belgrade, Yugoslavia.
- 1983. Diploma work in Institut for Biological Research Sinisa Stankovic, supervisor Dr Vojo Deretic.
- 1978-1983. B. Sci training, Molecular Biology and Physiology, major Biophysics, University of Belgrade, Yugoslavia.
Publications
Download-
2023Membrane platform protein PulF of the Klebsiella type II secretion system forms a trimeric ion channel essential for endopilus assembly and protein secretion., mBio 2023 Dec; (): e0142323.
-
2023The periplasmic coiled coil formed by the assembly platform proteins PulL and PulM is critical for function of the Klebsiella type II secretion system., Res Microbiol 2023 May; (): 104075.
-
2023Bacterial Type II Secretion System and Its Mitochondrial Counterpart., mBio 2023 Apr; 14(2): e0314522.
-
2023Structure and dynamic association of an assembly platform subcomplex of the bacterial type II secretion system., Structure 2023 Feb; 31(2): 152-165.e7.
-
20211 H, 15 N and 13 C resonance assignments of the C-terminal domain of PulL, a component of the Klebsiella oxytoca type II secretion system, Biomol NMR Assign . 2021 Oct;15(2):455-459. .
-
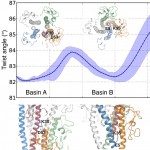
2021Computational and biochemical analysis of type IV pilus dynamics and stability., Structure 2021 Dec; 29(12): 1397-1409.e6.
-
20211H, 15 N and 13C resonance assignments of the C-terminal domain of PulL, a component of the Klebsiella oxytoca type II secretion system., Biomol NMR Assign 2021 Oct; 15(2): 455-459.
-
2021Analysis of diverse eukaryotes suggests the existence of an ancestral mitochondrial apparatus derived from the bacterial type II secretion system., Nat Commun 2021 May; 12(1): 2947.
-
2019Structure and function of minor pilins of type IV pili., Med Microbiol Immunol 2020 Jun; 209(3): 301-308.
-
2019Structure and Assembly of the Enterohemorrhagic Escherichia coli Type 4 Pilus., Structure 2019 Jul; 27(7): 1082-1093.e5.
-
+View full list of publications