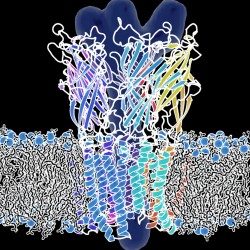
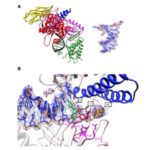
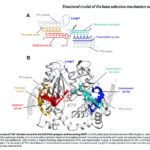
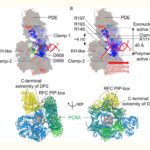
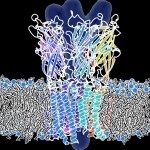
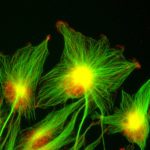
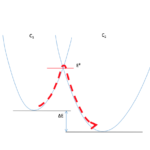
Un descriptif plus complet des activités de l’Unité se trouve sur http://lorentz.dynstr.pasteur.fr (en interne) ou bien (version externe) http://www.dynstr.pasteur.fr
Cliquez pour voir le graph
Connexions
Présentation
Membres

Marc Delarue
Responsable de Structure
Responsable
Groupes
Group
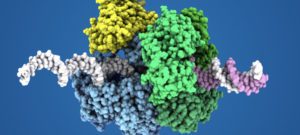
Groupe de Réplication de l’ADN

Ludovic Sauguet
Anciens Membres
2000
2000
Name
Position
2005
2006
Erik Lindahl
Post-doc
2005
2010
Félix Romain
Ingénieur
2005
2008
Jérémie Piton
PhD Student
2005
2008
Cyril Azuara
PhD Student
2008
2014
Jerome Gouge
PhD Student
2008
2012
Hugues Nury
Post-doc
2009
2010
Claudine Mayer
Prof. Paris Cité
2010
2010
Dennis Ptchelkine
Post-doc
2012
2016
Frederic Poitevin
PhD Student
2014
2020
Zaineb Fourati-Kammoun
Post-doc
2015
2020
Frédéric Poitevin
Post-doc
2015
2020
Filippo Romoli
Post-doc
2015
2020
Danièle Sinnaya
Administrative Staff
2015
2020
Irina Volahasina Randrianjatovo-Gbalou
Post-doc
2015
2020
Pierre Raia
PhD Student
2015
2020
Jérôme Loc’h
Post-doc
2015
2020
Haidai Hu
PhD Student
2015
2020
Victoire Prat
Undergraduate Student
2016
2020
Thomas Ybert
Post-doc
2017
2021
Dariusz Czernecki
PhD Student
2018
2022
Clement Madru
Post-doc
2018
2021
Camille Samson
Post-doc
2022
2024
Isciane Commenge
Ingénieur
2023
2023
Milena Milovanovic
Master Student