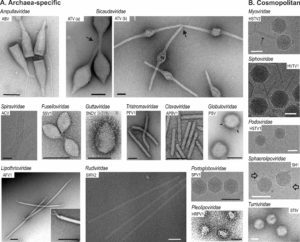
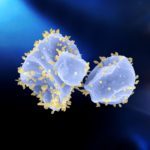
Research in the Archaeal Virology Unit focuses on addressing various fundamental questions at the interface of microbiology, virology and evolution: What molecular mechanisms allow viruses to subvert the host cells into virion-producing factories? What is the extent of viral diversity? What are the ultimate origins of different groups of viruses? How do these groups relate to each other as well as to other types of mobile genetic elements, such as plasmids and transposons? As a model for our research, we use viruses infecting archaea, which represent one of the least explored parts of the global virosphere. Archaea are hosts to viruses with some of the most unexpected virion morphologies, such as bottle-shaped, lemon-shaped, coil-shaped and droplet-shaped virions. The distinct nature of archaeal viruses also extends to their genomes, with the vast majority of the genes showing no similarity to sequences in the databases. We are exploring the extent of archaeal virus diversity in different habitats – from the human gut to hypersaline lakes and acidic hot springs – and study different aspects of their biology, including virion structure and assembly, molecular mechanisms of virus entry and egress as well as other aspects of virus-host interaction. During the past few years, we have isolated and characterized a number of new archaeal virus-host systems. This advancement provides us the opportunity to study not only virus-host interactions but also to explore different aspects of archaeal cell biology.