-
The bacterial capsule
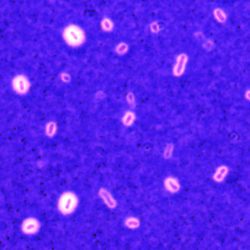
Our main axis of research is the study of the bacterial capsule. Extracellular capsules constitute the outermost layer of some bacteria and establish the first contact between the cell and its environment. Capsules are best known for their role in clinical settings, where they increase survival upon phagocytosis by eukaryotic cells and lower the sensitivity to antibiotics. They are thus considered a major virulence factor. However, capsules also play a critical role in the environment because they protect the cells from physical and chemical stresses. For example, they increase survival under desiccation and protect from antimicrobial peptides. They also enhance bacterial survival rates in mixed species communities and complex environments by, for instance, protecting bacteria from bacteriocins.
We aim to provide a global overview, across time scales, on the role of the capsule and how it affects the short, mid and long term genome evolution of a species. More specifically, using Klebsiella pneumoniae as a model system, a nosocomial pathogen causing lung, urinary and liver infections, our current research aims to:
- Quantify the short-term fitness and metabolic costs of capsule production.
- Understand how capsules affect bacterial adaptation to new environmentst hrough hundreds of generations.
- Determine the role of capsules in genetic transfer and the long-term evolution of genomes.
- Model the interplay between capsules and genetic exchange mechanisms, by building on an individual-based model (IBM), previously developed. This will allow us to make predictions about the capsule-related factors influencing genetic exchanges across cells and the evolutionary outcomes of multi-species communities under different environmental conditions. We can then test particular hypothesis in the lab.
-
Klebsiella prophages
Temperate phages have been shown to carry adaptive traits that increase the fitness of their bacterial host. Some of these traits are involved in virulence or bacterial survival under stress conditions. In the lab, we explore computationally the number and diversity of prophages across Klebsiella spp. and the factors that govern their occurrence and distribution. We are also interested in understanding the role that prophages can play in adaptation by assessing which bacterial traits they encode, with a focus on antibiotic resistance genes. Finally, we experimentally validate computational predictions, for instance, we study the ability of these prophages to excise, infect and lysogenize other Klebsiella strains.
-
Bacterial co-evolution
We are also interested in the social interactions taking place within communities. We want to determine to what extent phylogenetic distance between interactants influence total group productivity and the emergence of competitive or cooperative interactions. By using a novel protocol of single-cell digital PCR, we determine population dynamics over hundreds of generations across different environments relevant to Klebsiella spp tropism.