About
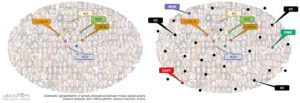
Although mouse models have been invaluable to study human diseases, classical mouse laboratory strains (figure, left panel, colored dots) present much smaller genetic diversity than the human population, limiting the range of phenotypic variations which can be observed in mice. A consortium of mouse geneticists and laboratories (USA, Israel, Australia) have developed a new collection of inbred strains which result from the crossing between eight founder strains, three of which were established from different mouse subspecies. This resource provides access to a range of genetic diversity similar to that of the human population (right panel, black dots). As a result, the CC allows the discovery of new phenotypes and the identification of their genetic control.
The central animal facility has imported and maintains 35 CC strains under the scientific supervision of the Mouse Genetics Laboratory. These strains are used in a growing number of projects campus-wide and in external collaborations. We are using them to study host genetic control of infections with Salmonella and Zika virus.
A “CC-Club” was created in 2017 as a forum where researchers can discuss concepts, ideas, praticalities, study design, results and grant applications.
A joint project with the Institut Pasteur Bioinformatics Hub aims at developing a database to store CC experimental data and a user-friendly interface to perform QTL mapping.