Présentation
Recipient: Gerald Spaeth, coordinator
Agency: Campus France, MEAE, Type: Bi-national program France/Israel, 04/2018 – 03/2020
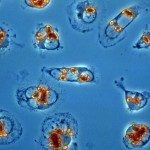
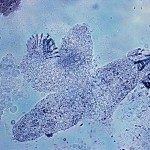
Leishmaniasis is among the five most serious parasitic diseases worldwide. The cutaneous form of the disease is prevalent in Israel with a sharp rise in the last years, while the fatal visceral form is emerging in the EU. The therapy is very poor especially for Leishmania tropica, which is endemic in Israel. Resistance to common drugs also developed in the last years. There is thus an urgent need for new anti-leishmanial treatments. An excellent target for chemotherapy is represented by the Leishmania ribosome, which in contrast to the mammalian ribosome undergoes extensive rRNA fragmentation mediated by highly abundant and trypanosomatid-specific small nucleolar RNAs (snoRNA), a phenomenon discovered by the Israeli group. Our proposal combines the complementary expertise of the French and Israeli partners in experimental evolution and RNA biology to model snoRNA selection in the field and in culture and to analyze their unique biological functions using genomics approaches, with the final aim to validate Leishmania non-coding RNAs and ribosomal biology as novel targets for anti-leishmanial intervention.
Our main hypothesis is that small nucleolar RNAs, which direct processing and modification on rRNA, are amplified during selection for parasite fitness in changing environmental conditions. The French group developed over the years an in vitro evolution system to study genomic mechanisms of parasite fitness and demonstrated that parasites adapt to in vitro culture through an evolutionary process by amplification of specific chromosomes that are enriched in specific snoRNAs. The Israeli group described all the Leishmania snoRNA and elucidate their function. We hypothesize that snoRNA amplification is a dominant feature of the adaptation because it may enhance processing and produce hypermodified rRNA thus generating ribosomes that are better adjusted to withstand stress. In the proposal we present three objectives (1) to apply computational pipelines established in the partner teams to model the evolutionary history of snoRNAs across the Leishmania genus, and to model snoRNA selection in the field, (2) to correlate Leishmania genome dynamics during evolutionary adaptation to changes in RNA biology and phenotype, and (3) to apply functional genomics approaches to map and correlate changes in RNA modification (Nms and Ψ) with the levels of endogenous and transgenic snoRNA expression. We will establish the computational framework to model and quantify the impact of snoRNA transgenic expression on parasite post-transcriptional regulation and gene expression. Our multi-disciplinary approach directly addresses a major public health problem in Israel and in France, and will drive scientific discovery on Leishmania RNA biology, gene regulation and genome instability, with important translational potential for innovative, anti-parasitic intervention strategies.