About
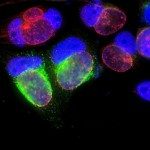
The membrane of the inclusion is key for the regulation of the exchanges between the bacteria and the host. We study its biogenesis and how metabolites from the host are transported to the lumen of the inclusion.
Identification of type III secreted proteins and functional studies
Throughout their cycle in the host cell, chlamydiae remain in a compartment called an inclusion. They use a type III secretion mechanism to translocate proteins into the host cell. These proteins allow the bacteria to manipulate host activities to their benefit. By studying the function of these proteins we aim at understanding how the bacteria escape innate defense from the host and thrive in the vacuolar environment.
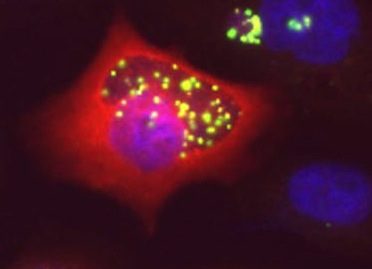
Living on host resources
Chlamydiae uses host metabolites for growth. They lack enzymes for several biosynthetic pathways, and exploit their host to compensate. We study the mechanisms by which cytosolic metabolites reach the inclusion lumen, and the consequences of the infection on the metabolism of the host.