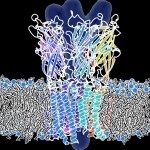
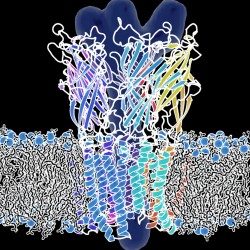
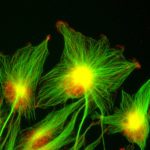
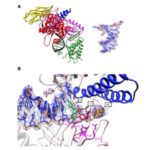
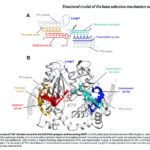
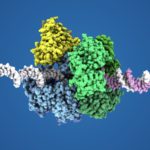
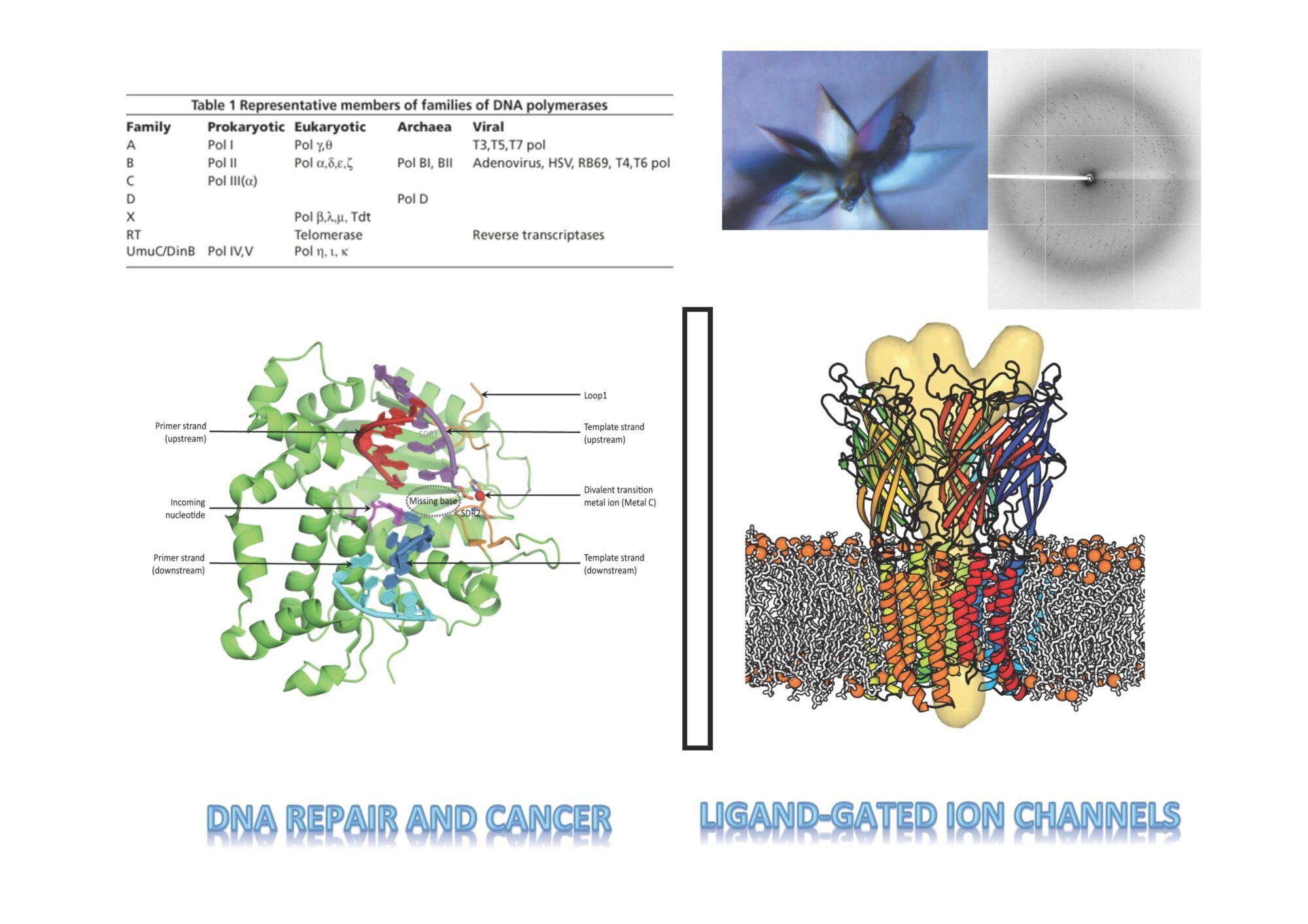
- We use biophysical experimental techniques such as crystallography and cryo-EM to visualize at the atomic level the structure of molecules essential to life, such as DNA polymerases involved in DNA Repair and Cancer and ion channels involved in electric nerve signalling (cell-cell communications).
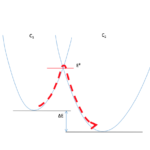
- We complement them with molecular and normal modes dynamics, so as to go beyond the essentially static pictures given by these methods.
- We also try to better understand the electrostatics of macromolecules and their interaction with the solvent and ligands, in order to be able to predict their binding properties.
- Our main goal is to design structure-inspired drugs (pharmacology) and re-design active site(s) to make them accept other substrates (synthetic biology).
- More details can be found at http://dynstr.pasteur.fr or (internally) http:/lorentz.dynstr.pasteur.fr
Click to view graph
Connections
Click to view timeline
Timeline
About
Transversal Projects
Projects
Software
Publications
Download-
2024MinActionPath2: path generation between different conformations of large macromolecular assemblies by action minimization., Nucleic Acids Res 2024 May; (): .
-
2023Molecular basis for proofreading by the unique exonuclease domain of Family-D DNA polymerases., Nat Commun 2023 Dec; 14(1): 8306.
-
2023Analyzing the Geometry and Dynamics of Viral Structures: A Review of Computational Approaches Based on Alpha Shape Theory, Normal Mode Analysis, and Poisson–Boltzmann Theories, Viruses 2023, 15, 1366.
-
2023DNA-binding mechanism and evolution of replication protein A., Nat Commun 2023 Apr; 14(1): 2326.
-
2023Reclassification of family A DNA polymerases reveals novel functional subfamilies and distinctive structural features, Nucleic Acids Research.
-
2023Computing the Gromov-Wasserstein Distance between Two Surface Meshes Using Optimal Transport, Algorithms 2023, 16, 131.
-
2021Structural dynamics and determinants of 2-aminoadenine specificity in DNA polymerase DpoZ of vibriophage ϕVC8., Nucleic Acids Res 2021 Nov; 49(20): 11974-11985.
-
2021Extracting Dynamical Correlations and Identifying Key Residues for Allosteric Communication in Proteins by correlationplus., J Chem Inf Model 2021 Oct; 61(10): 4832-4838.
-
2021Characterization of a triad of genes in cyanophage S-2L sufficient to replace adenine by 2-aminoadenine in bacterial DNA., Nat Commun 2021 08; 12(1): 4710.
-
2021Parameterizing elastic network models to capture the dynamics of proteins., J Comput Chem. 2021 Jun 11..
-
+View full list of publications