Présentation
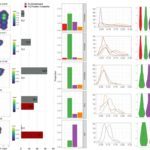
Listeriomics integrates all complete genomes, transcriptomes and proteomes published for Listeria species to date. It allows navigating among all these datasets with enriched metadata in a user-friendly format.
The main purpose of the website is to give scientists a quick and easy access to tools created for answering the four questions one has when starting a study on a specific genomic element:
- What are the known functions of a genomic feature in a given strain and homologies in closely related strains?
- What are the biological conditions in which a genomic feature is transcribed?
- What are the biological conditions in which a genomic feature is translated?
- What is the regulation network involved with a genomic feature of interest?
The website integrates different type of tools for “omics” data analyses:
- A highly interactive genome viewer for display of gene expression arrays, tiling arrays, and RNA-Seq datasets along with proteomics and genomics datasets.
- Expression and protein atlas, which are tools to connect every genomic feature (genes, smallRNAs, antisenseRNAs) or proteins to the most relevant “omics” data.
- A tool to explore gene conservation throughout Listeria strains.
- A co-expression network tool to discover new potential regulation.
Reference : Bécavin et al., mSystems 2017