Link to Pubmed [PMID] – 25048412
FEMS Microbiol. Lett. 2014 Sep;358(1):1-10
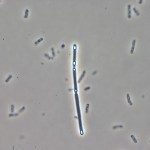
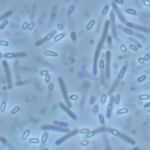
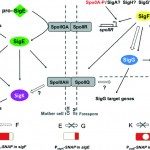
Clostridium difficile, a Gram-positive, anaerobic, spore-forming bacterium, is a major cause of nosocomial infections such as antibiotic-associated diarrhea. Spores are the vector of its transmission and persistence in the environment. Despite the importance of spores in the infectious cycle of C. difficile, little was known until recently about the control of spore development in this enteropathogen. In this review, we describe recent advances in our understanding of the regulatory network controlling C. difficile sporulation. The comparison with the model organism Bacillus subtilis highlights major differences in the signaling pathways between the forespore and the mother cell and a weaker connection between morphogenesis and gene expression. Indeed, the activation of the SigE regulon in the mother cell is partially independent of SigF although the forespore protein SpoIIR, itself partially independent of SigF, is essential for pro-SigE processing. Furthermore, SigG activity is not strictly dependent on SigE. Finally, SigG is dispensable for SigK activation in agreement with the absence of a pro-SigK sequence. The excision of the C. difficile skin element is also involved in the regulation of SigK activity. The C. difficile sporulation process might be a simpler, more ancestral version of the program characterized for B. subtilis.