About
In comparison to other bacterial species, genetics and molecular basis of the pathogenesis of leptospires are in their infancy. An improved understanding of leptospiral pathogenesis requires reliable genetic tools for functional genetic analysis of Leptospira spp.
We generated kanamycin, gentamycin, and spectinomycin resistance cassettes as selectables markers for genetic manipulations in Leptospira spp. We have shown that the E. coli lac system is functional in Leptospira and can be used to control the expression of genes. The Photorhabdus luminescens luxCDABE cassette have been transferred into Leptospira strains to produce luminescent leptospires in live mice. These bioluminescent leptospires are novel tools that will be useful to test the efficacy of antiobiotic treatments or vaccines against leptospirosis. We have previously generated replicative vectors for the saprophyte L. biflexa but none of these plasmids showed autonomous replication in pathogenic strain. In a recent study, cloning of the putative 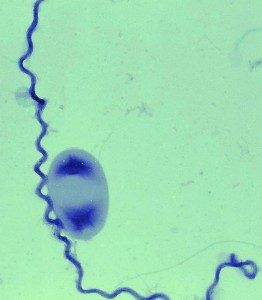
The genus Leptospira is composed of both saprophytic and pathogenic members, such as L. biflexa and L. interrogans, respectively. L. biflexa is a “fast-growing” organism with a generation time of 5 hours, in comparison to 20 hours for pathogenic strains. Our group has sequenced the genome of the saprophyte L. biflexa and we developed a number of tools for genetic manipulation of L. biflexa. These advances provide us with an opportunity to use L. biflexa as a model bacterium for studying leptospiral biology. We also showed that L. biflexa can serve as a surrogate host to characterize the role of key virulence factors of the causative agent of leptospirosis. In contrast to the pathogens that are poorly transformable, thousands of random mutants can be readily obtained in L. biflexa by random insertion of Himar1 in the genome, thereby generating extensive libraries of mutants that could be screened for phenotypes affecting diverse aspects of the biology of the bacterium.