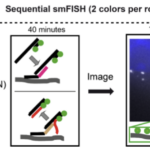
I’m a trained biophysicist and develop integrated approaches combining fluorescence imaging, wetlab experiments, computerised image analysis, and mathematical modeling. We apply this methods to study RNA regulation across several scales, ranging from sub-cellular RNA localization to cell-type mapping in tissue.
I’m currently heading the Quantitative RNA imaging group.