Présentation
Going the extra mile in Gene Set Enrichment analysis: seeing the data with g-RITA
Gene Set Enrichment Analysis (GSEA) is a powerful approach for interpreting OMIC data based
on the functional annotation of genes. Such methods are useful for identifying enriched gene sets,
i.e.,groups of related genes involved in the same molecular pathways, biological processes, cellular
compartments, or in other common features.The resulting enriched gene sets provide information on
the biological functions and processes associated with specific conditions.GSEA tools share, overall, the same output format of i) a list of gene sets enriched in a given biological condition, ii) the number of genes in the dataset belonging each gene set, and iii) the p-value associated with the test statistic. This format fails to reveal the relationship among the listed gene sets.
Adittionaly, GSEA methods do not usually provide graphical representations of the results.
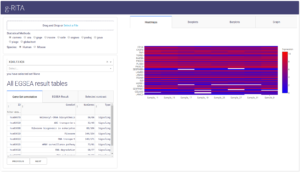
g-RITA (gene enRichment InteracTive visuAlisation) is an interactive web interface to visualise
and further explore the results for functional analysis of RNAseq data. It combines i) gene-set level and ii) gene level information in a single analysis allowing the analysis of results at different granularity levels, which largely improves the interpretation of the underlying molecular mechanisms of the data. g-RITA GSEA results are based on the EGSEA R package.
g-RITA interface
More about EGSEA R package