Présentation
One of the major challenges of modern medicine is to understand how the human organism defends itself against invasions and diseases. The biggest mystery is the human immune system, and understanding this ultimately requires knowledge of the sequence repertoire of human antibodies (Ab) and their respective antigens. Current state-of-art Ab analysis, based on an enzymatic digestion, destroys information on the connectivity of heavy and light chains and is not able to achieve an extensive sequencing of complementary determining regions (CDRs) that are binding to antigens. The solution is therefore to keep proteins intact before mass spectrometry analysis (MS) and perform sequencing in the gas phase (top-down MS/MS approach). However complete sequencing of intact proteins larger than 20 kDa still represents an insurmountable challenge to MS.
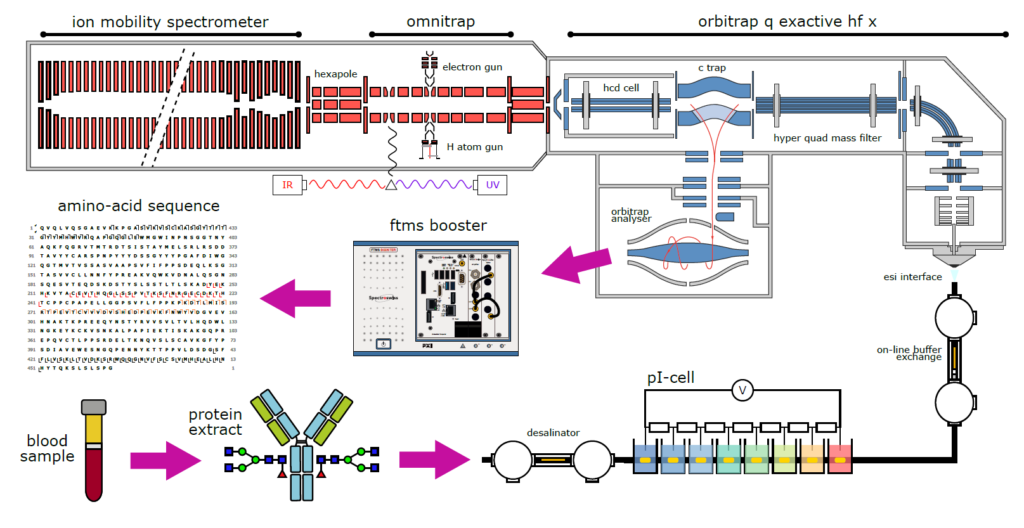
The TopSpec project is aiming at developing cutting-edge high-resolution mass spectrometry approaches for the analysis of intact antibodies and antibody-based drugs. The core of the project is the development of a novel top-down tandem mass spectrometry (MS/MS) platform. One major innovation is the implementation of novel radical gas-phase ion-electron and ion-atom reactions to better sequence large intact proteins. TopSpec will also pioneer the deconvolution of massively overlapping isotopic clusters, solving one of the greatest challenges in top-down MS/MS of large molecules. This technological breakthrough will enable scientists from academia and industry to better characterize antibodies, enhancing therefore our ability to explore quickly new effective therapies against major global health issues.
The TopSpec project is led by a consortium of European key experts in ultrahigh-resolution mass spectrometry, top-down proteomics and MS data processing (Karolinska Institutet, Fasmatech, Thermo Fisher Scientific, Spectroswiss, Biomotif, The Nottingham Trent University, Institut Pasteur and MS Vision) and has received funding from the European Horizon 2020 research and innovation program.