Présentation
Jean-Pierre Levraud, Valerio Laghi, Pedro Hernandez-Cerda, Valérie Briolat, Nuno Palha, Gabriella Passoni
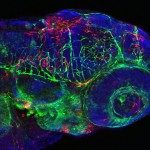
Viruses are ubiquitous but elusive pathogens, invisible to the light microscope. How do our host defenses counter their spread within our organisms is only known in very general terms. The transparency of the zebrafish larva, combined with appropriate microscopic and genetic tools, allows us to adress this question at the whole organism level and single-cell resolution.
Anti-viral innate defenses
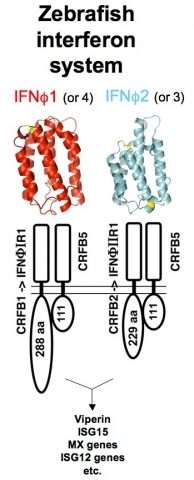
Being vertebrates, zebrafish possess immune defenses that are similar to our own, but still incompletely characterized as compared to other model animals. We tackle this issue by functional, genomic and transcriptomic analyses of relevant genes. Interferons (IFNs) orchestrate innate antiviral responses. These cytokines are secreted when cells detect viruses; in response, cells harboring IFN receptors express many genes (ISGs, for IFN-stimulated genes) and acquire a virus-resistant state. In the past years, we have identified by genetic approaches the receptors of zebrafish IFNs (Levraud et al., 2007 ; Aggad et al., 2009). Fish viro-induced IFNs fall into two subgroups with low sequence homology, and with distinct receptors ; however, we found by crystallographic analysis that they share a similar 3D fold, matching that of mammalian type I IFNs (Hamming et al., 2011). We have also characterized a number of fish ISGs. Unexpectedly, this led us to characterize three highly expanded families of TRIM (TRIpartite Motif) genes which highly diversified C-terminal sequences, suggesting a role in recognition of variable ligands (van der Aa et al., 2009 ; Boudinot et al., 2011). We hypothesize that this diversification may have been driven by interactions with natural zebrafish viruses, which would be restricted by these TRIM proteins.
IHNV : a virus causing a fatal hemorrhagic disease
Althought no natural zebrafish virus has been characterized so far, we and other labs have established a few models of experimental viral infection of the zebrafish. We have performed a detailed analysis of the infection with IHNV25, a strain of Infectious Hematopoietic Necrosis Virus, a salmonid rhabdovirus adapted to growth up to 25°C. Using various techniques, we could follow the progression of the infection in whole larvae (Ludwig et al., 2011). Vascular endothelial cells are the primary targets of the virus, resulting in hemorrhages ; infection then spreads to other tissues. Although IFN could protect the larva if provided early, the host response comes too late and the outcome of the infection is invariably fatal in unmanipulated larvae.
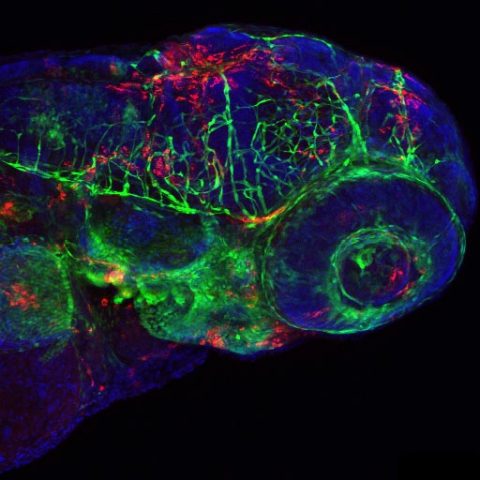
Confocal image of a fli1:GFP larva inoculated 24h before with IHNV25. Endothelial cells appear in green, infected cells in red, nuclei in blue. Image by Nuno Palha.
Chikungunya virus : a human pathogen that elicits a protective interferon response
More recently, we have developed a new model of infection with a human mosquito-borne virus, Chikungunya virus (ChikV), considered a re-emerging pathogen. ChikV can replicate efficiently in zebrafish larvae but the infection recedes spontaneously after a few days. We have shown that this is due to the endogenous interferon response (Palha et al., in preparation). Using a GFP recombinant virus and a zebrafish transgenic RFP reporter under the control of an IFN promoter, we can follow in real-time the progression of the infection and the responding host cells.
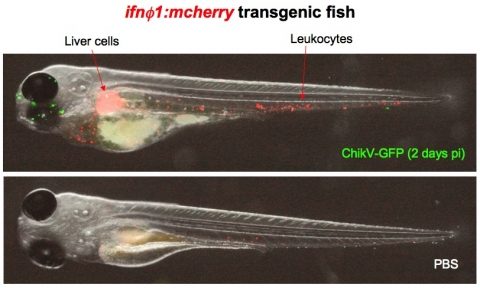
Simultaneous live imaging of the viral infection and the host response with transgenic zebrafish: interferon-producing cells in ChikV-GFP-infected larvae
We are affiliated to the Fishforpharma ITN network. Collaborations Pierre Boudinot, INRA, Jouy-en-Josas, France Georges Lutfalla, CNRS, Montpellier, France Rune Hartmann, Université d’Aarhus, Danemark Olivier Schwartz, Institut Pasteur, Département de Virologie, Institut Pasteur, France