Cliquez pour voir la ligne de temps
Ligne de temps
Projets
Outils
Anciennes Équipes
Publications
Télécharger-
2024Profiling Cullin4-E3 ligases interactomes and their rewiring in influenza A virus infection., Mol Cell Proteomics 2024 Oct; (): 100856.
-
2022SKAP2 modular organization differently recognizes SRC kinases depending on their activation status and localization., Mol Cell Proteomics 2022 Nov; (): 100451.
-
2022A proteome-scale map of the SARS-CoV-2-human contactome., Nat Biotechnol 2022 Oct; (): .
-
2018The SRC-family tyrosine kinase HCK shapes the landscape of SKAP2 interactome, Oncotarget 2018 Mar;9(17):13102-13115.
-
2017Inhibition of the inflammatory response to stress by targeting interaction between PKR and its cellular activator PACT, Sci Rep. 2017 Nov 23;7(1):16129.
-
2015Quantifying domain-ligand affinities and specificities by high-throughput holdup assay, Nat. Methods 2015 Aug;12(8):787-93.
-
2013A comparative approach to characterize the landscape of host-pathogen protein-protein interactions, J Vis Exp 2013;(77):e50404.
-
2012Comparative analysis of virus-host interactomes with a mammalian high-throughput protein complementation assay based on Gaussia princeps luciferase, Methods 2012 Dec;58(4):349-59.
-
2012Mapping of Chikungunya virus interactions with host proteins identified nsP2 as a highly connected viral component., J Virol 2012 Mar; 86(6): 3121-34.
-
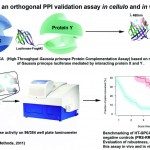
2011Benchmarking a luciferase complementation assay for detecting protein complexes, Nat. Methods 2011 Dec;8(12):990-2.
-
+Voir la liste complète de publications