Cliquez pour voir le graph
Connexions
Cliquez pour voir la ligne de temps
Ligne de temps
Projets Transversaux
Projets
Logiciel
Outils
Brevets
Anciennes Équipes
Publications
Télécharger-
2024Structure and Dynamics of Type 4a Pili and Type 2 Secretion System Endopili, In: Macromolecular Protein Complexes V, vol 104. Chapter 21. p549-563.
-
2024A comprehensive dataset of protein-protein interactions and ligand binding pockets for advancing drug discovery, Scientific Data , 2024, 11 (1), pp.402. ⟨10.1038/s41597-024-03233-z⟩.
-
2023Structure and dynamic association of an assembly platform subcomplex of the bacterial type II secretion system., Structure 2023 Feb; 31(2): 152-165.e7.
-
2022Building Protein Atomic Models from Cryo-EM Density Maps and Residue Co-Evolution., Biomolecules 2022 Sep; 12(9): .
-
2022Secondary structure and 1H, 15 N & 13C resonance assignments of the periplasmic domain of OutG, major pseudopilin from Dickeya dadantii type II secretion system., Biomol NMR Assign 2022 Apr; (): .
-
2022Bat coronaviruses related to SARS-CoV-2 and infectious for human cells., Nature 2022 Apr; 604(7905): 330-336.
-
2021InDeep: 3D fully convolutional neural networks to assist in silico drug design on protein-protein interactions., Bioinformatics 2022 Feb; 38(5): 1261-1268.
-
20211 H, 15 N and 13 C resonance assignments of the C-terminal domain of PulL, a component of the Klebsiella oxytoca type II secretion system, Biomol NMR Assign . 2021 Oct;15(2):455-459. .
-
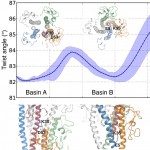
2021Computational and biochemical analysis of type IV pilus dynamics and stability., Structure 2021 Dec; 29(12): 1397-1409.e6.
-
2021Host-Pathogen Adhesion as the Basis of Innovative Diagnostics for Emerging Pathogens., Diagnostics (Basel) 2021 Jul; 11(7): .
-
+Voir la liste complète de publications