Link to Pubmed [PMID] – 29150513
Appl. Environ. Microbiol. 2017 Nov;
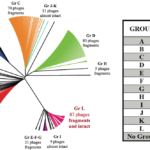
Clostridioides difficile (formerly Clostridium difficile) is a pathogenic bacterium displaying great genetic diversity. A significant proportion of this diversity is due to the presence of integrated prophages. Here, we provide an in-depth analysis of phiCD211, also known as phiCDIF1296T, the largest phage identified in C. difficile so far, with a genome of 131-kbp. It shares morphological and genomic similarity with other large siphophages like phage 949 infecting Lactococcus lactis and phage c-st infecting Clostridium botulinum. A PhageTerm analysis indicated the presence of 378-bp direct terminal repeats at the phiCD211 genome termini. Among striking features of phiCD211, the presence of several transposase and integrase genes suggests past recombination events with other mobile genetic elements. Several gene products potentially influence the bacterial lifestyle and fitness, including a putative AcrB/AcrD/AcrF multidrug resistance protein, an EzrA septation ring formation regulator, and a spore protease. We also identified a CRISPR locus and a cas3 gene. We screened 2,584 C. difficile genomes available and detected 149 prophages sharing ≥80% nucleotide identity with phiCD211 (5% prevalence). Overall, phiCD211-like phages were detected in C. difficile strains corresponding to 21 different MLST groups, showing their high prevalence. Comparative genomic analyses revealed the existence of several clusters of highly similar phiCD211-like phages. Of note, large chromosome inversions were observed in some members, as well as multiple gene insertions and module exchanges. This highlights the great plasticity and gene coding potential of the phiCD211/phiCDIF1296T genome. Our analyses also suggest active evolution involving recombination with other mobile genetic elements.IMPORTANCEClostridioides difficile is a clinically important pathogen representing a serious threat to human health. Our hypothesis is that genetic differences between strains caused by the presence of integrated prophages could possibly explain the apparent differences observed in the virulence of different C. difficile strains. In this study, we provide a full characterization of phiCD211, also known as phiCDIF1296T, the largest phage known to infect C. difficile so far. Screening 2,584 C. difficile genomes revealed the presence of highly similar phiCD211-like phages in 5% of the strains analyzed, showing their high prevalence. Multiple genome comparisons suggest that evolution of the phiCD211-like phage community is dynamic, and some members have acquired genes that could possibly influence bacterial biology and fitness. Our study further supports the relevance of studying phages in C. difficile to better understand the epidemiology of this clinically important human pathogen.