Link to Pubmed [PMID] – 32022687
Link to DOI – 10.7554/eLife.51247
Elife 2020 Feb; 9():
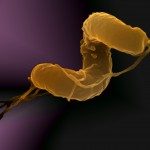
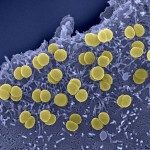
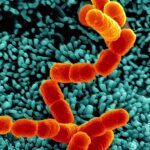
Lytic transglycosylases (LT) are enzymes involved in peptidoglycan (PG) remodeling. However, their contribution to cell-wall-modifying complexes and their potential as antimicrobial drug targets remains unclear. Here, we determined a high-resolution structure of the LT, an outer membrane lipoprotein from Neisseria species with a disordered active site helix (alpha helix 30). We show that deletion of the conserved alpha-helix 30 interferes with the integrity of the cell wall, disrupts cell division, cell separation, and impairs the fitness of the human pathogen Neisseria meningitidis during infection. Additionally, deletion of alpha-helix 30 results in hyperacetylated PG, suggesting this LtgA variant affects the function of the PG de-O-acetylase (Ape 1). Our study revealed that Ape 1 requires LtgA for optimal function, demonstrating that LTs can modulate the activity of their protein-binding partner. We show that targeting specific domains in LTs can be lethal, which opens the possibility that LTs are useful drug-targets.Bacteria are surrounded by a tough yet flexible wall that protects the cell and serves as an anchor for several of the cell’s structures. This cell wall contains a large mesh-like molecule called peptidoglycan made of many repeated building blocks. When a bacterial cell divides in two, it needs to make more of this material. Making peptidoglycan involves two different sets of enzymes working together: “polymerases” are the enzymes that link the individual building blocks to peptidoglycan, one after the other; while “lytic transglycosylases” are enzymes that modify the peptidoglycan to create space for the addition of new building blocks and for assemblies of proteins that must span the cell wall. Lytic transglycosylases are known to assemble with other proteins and enzymes to form the cell’s peptidoglycan-modifying machinery, but it was not clear exactly what purpose they serve within these “enzyme complexes”. It was also unclear whether these enzymes would be good targets for new antibiotics. To help answer these questions, Williams et al. looked at a lytic transglycoslyase called LtgA. This enzyme is originally from Neisseria meningitidis, a bacterium that can cause meningitis and life-threatening sepsis in humans. Williams et al. discovered that part of the enzyme’s active site – the region of an enzyme where the chemical reaction takes – can switch from an ordered helix to a disordered, flexible loop. Bacteria were then genetically engineered to make a version of the enzyme that lacked this helix. These bacteria had weaker cell walls and were deformed; they were also less able to grow and divide, both in the laboratory and in a mouse model of infection. Further analysis showed that the deletion of the helix from the enzyme resulted in the peptidoglycan being modified much more than normal, which could likely explain their reduced virulence. Williams et al. also found that deleting the helix from LtgA interfered with the activity of a protein that interacts with this enzyme, called Ape1, which also contributed to the fragility of the cell wall. This shows that lytic transglycosylases assembled into enzyme complexes can alter the activities of other proteins in the complex. Together these findings show that researchers could target one enzyme in a complex in bacteria, and disrupt the activity of other proteins in that complex. This highlights the possibility of considering enzyme complexes as useful targets for new drugs, which is important considering the current problem of antibiotic resistance.