About
Two-component regulatory proteins function in bacteria as sensory and adaptive factors in response to a wide range of environmental stimuli. Some two-component systems, such as PhoP/PhoQ, control the transcription of key virulence genes in several intracellular bacterial pathogens, including Salmonella spp., Shigella spp. and Yersinia spp. In collaboration with Carlos Martín’s laboratory (University of Zaragoza, Spain), we have constructed, by allelic replacement, phoP and phoQ mutants of several M. tuberculosis strains. These mutants are impaired for in vivo multiplication. These mutants are not only attenuated, but also protect against a challenge with M. tuberculosis wild type strains. The particular morphological and cytochemical properties of phoP and phoP-phoR knock-out mutants of M. tuberculosis then led us to investigate the lipid content of these strains. Our results indicated that PhoP coordinately and positively regulates the synthesis of methyl-branched fatty acid-containing acyltrehaloses known to be restricted to the pathogenic species of the M. tuberculosis complex, namely diacyltrehaloses, polyacyltrehaloses, and sulfolipids. The analysis of transcriptional profiles in phoP mutants has evidenced a complex network of regulation including many genes involved in lipid biosynthesis. More in depth genetic studies will define what are the genes that are directly regulated by PhoP/PhoQ. Similarities between M. tuberculosis phoP-phoR mutants and the attenuated laboratory strain M. tuberculosis H37Ra in terms of morphological and cytochemical properties, lipid content, gene expression and virulence attenuation then prompted us to analyze the functionality of this two-component regulator in the latter strain. Sequence analysis revealed a base substitution resulting in a one amino acid change in the likely DNA-binding region of PhoP in H37Ra relative to H37Rv. Using gel-shift assays, and in collaboration with Deshmuk Gopaul’s team (Institut Pasteur), we then showed that this mutation abrogated the ability of the H37Ra PhoP protein to bind to a 40-bp segment of its own promoter. Complementation of H37Ra with phoP from H37Rv fully restored sulfolipid, diacyltrehalose and polyacyltrehalose synthesis clearly indicating that the lack of production of these lipids in H37Ra is solely due to the point mutation in phoP. Finally, using a pks2-3/4 knock-out mutant of M. tuberculosis H37Rv, evidence was further provided that the above mentioned polyketide-derived acyltrehaloses do not significantly contribute to the virulence of the tubercle bacillus in a mouse model of infection. Reasons for the attenuation of H37Ra thus most likely stand in other virulence factors, many of which are expected to belong to the PhoP regulon and another of which, unrelated to PhoP, appears to be the lack of production of DIM in this strain.
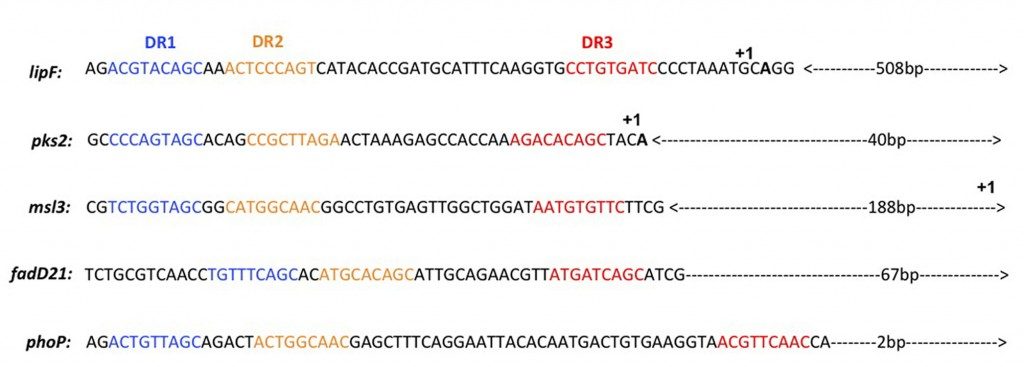
A consensus sequence derived from the analysis of PhoP binding to four gene promoter region were derived from our binding assays analyzis. We show for the first time the involvement of a third direct repeat motif in this binding reaction. The consensus was used to study the global regulation mediated by PhoP in M. tuberculosis. This allowed to identify the putative M. tuberculosis PhoP regulon.
Figure 1 : Identification of the DR1, DR2 and DR3 repeats in the upstream regions of the phoP, lipF, pks2, msl3 and fadD21 genes.
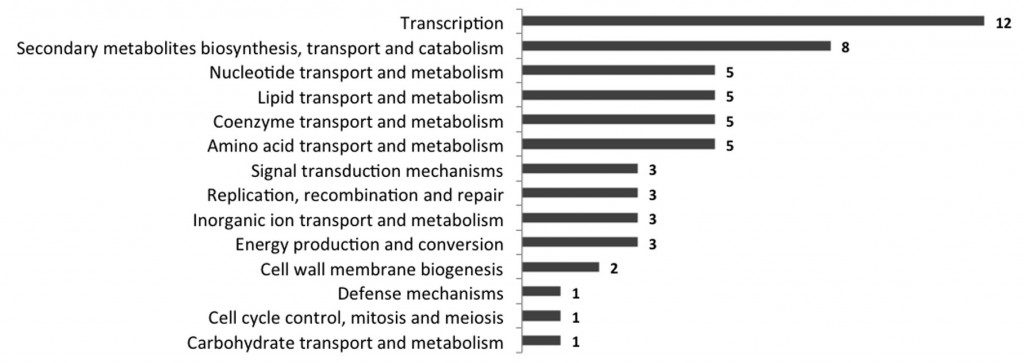
Figure 2 : Distribution of the identified consensus sequences of DR1, DR2 and DR3 in the upstream regions of all Mycobacterium tuberculosis H37Rv genes (according to the Clusters of Orthologous Groups (COG) classification).