Click to view graph
Connections
Click to view timeline
Timeline
Projects
Former Teams
Publications
Download-
2019Paracoccus haematequi sp. nov., isolated from horse blood., Int J Syst Evol Microbiol 2019 Jun; 69(6): 1682-1688.
-
2019Pigmentiphaga humi sp. nov., isolated from soil amended with humic acid., Int J Syst Evol Microbiol 2019 Jun; 69(6): 1573-1578.
-
2019Filibacter tadaridae sp. nov., isolated from within a guano pile from a colony of Mexican free-tailed bats Tadarida brasiliensis., Int J Syst Evol Microbiol 2019 May; 69(5): 1438-1442.
-
2019Implementation of the Nagoya Protocol within the Collection of Institut Pasteur, Access Microbiol. 2019 Jan doi: 10.1099/acmi.0.000008 .
-
2018Isolation by Miniaturized Culture Chip of an Antarctic bacterium sp. with antimicrobial and anthelmintic activity, Biotechnol Rep (Amst) 2018 Dec;20:e00281.
-
2018Draft Genome Sequence of the Fish Pathogen Flavobacterium columnare Genomovar III Strain PH-97028 (=CIP 109753)., Genome Announc 2018 Apr; 6(14): .
-
2018Revisiting the taxonomy of the genus Elizabethkingia using whole-genome sequencing, optical mapping, and MALDI-TOF, along with proposal of three novel Elizabethkingia species: Elizabethkingia bruuniana sp. nov., Elizabethkingia ursingii sp. nov., and Elizabethkingia occulta sp. nov., Antonie Van Leeuwenhoek 2018 Jan; 111(1): 55-72.
-
2017Psychrobacter pasteurii and Psychrobacter piechaudii sp. nov., two novel species within the genus Psychrobacter., Int J Syst Evol Microbiol 2017 Sep; 67(9): 3192-3197.
-
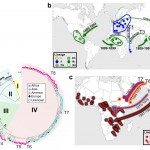
2017Global and regional dissemination and evolution of Burkholderia pseudomallei, Nat Microbiol. 2017 Jan 23;2:16263. doi: 10.1038/nmicrobiol.2016.263.
-
2016Whole-Genome Sequence of Corynebacterium auriscanis Strain CIP 106629 Isolated from a Dog with Bilateral Otitis from the United Kingdom., Genome Announc 2016 Aug; 4(4): .
-
+View full list of publications