Présentation
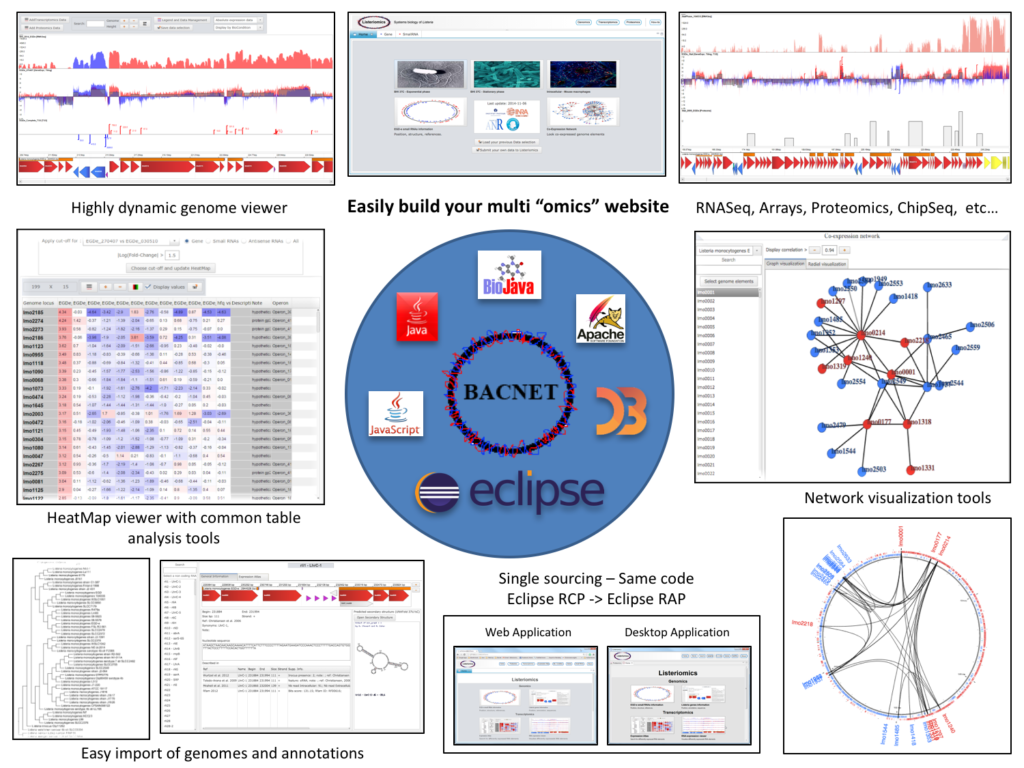
BACNET is a Java based platform to develop website for multi-omics analysis. Our platform integrates different type of tools, the three most important being:
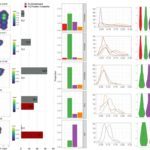
A table tools for Heatmap drawing and manipulation.
Genome viewer for displaying gene expression array (DNA microarray), tiling array, and RNA-Seq data along with proteomics and genomics data. This genome viewer is able to load quickly every data using streaming techniques, to zoom through the data from a nucleotide scale to a genome-wide scale, and overlay heterogeneous “omics” data.
An expression atlas, inspired from EBI Expression Atlas, for easily design a query based tool which connects every genomics elements (genes, smallRNAs, antisenseRNAs) to the most relevant “omics” data.
BACNET allows also to develop local software and websites with the same code. To reach this goal we used two highly connected APis , Eclipse RCP and Eclipse RAP (single sourcing).
A referent website for Listeria species called Listeriomics has been developed using BACNET platform. The main purpose of the Listeriomics website is to give scientists a quick and easy access to tools created for answering the four questions one has when starting a study on a specific genomic element:
What are the known functions of a genomic feature in a given strain and homologies in closely related strains?
What are the biological conditions in which a genomic feature is transcribed?
What are the biological conditions in which a genomic feature is translated?
What is the regulation network involved with a genomic feature of interest?
The following tutorials will show you how to build such a website using BACNET platform: https://gitlab.pasteur.fr/bacnet/Bacnet-public/wikis/home