Présentation
Transmission electron microscopy
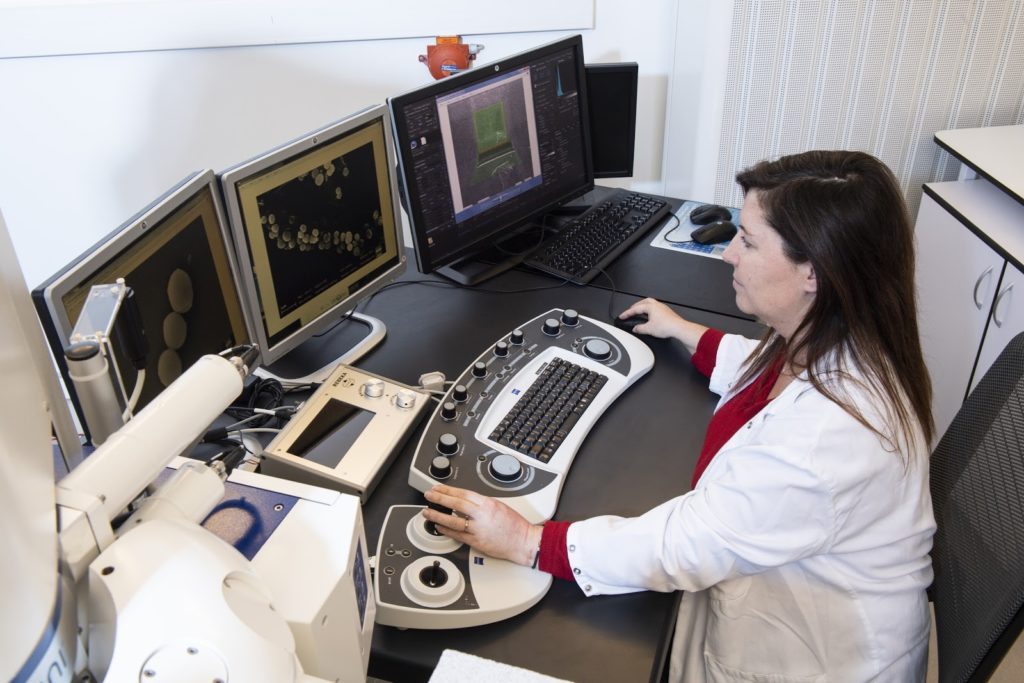
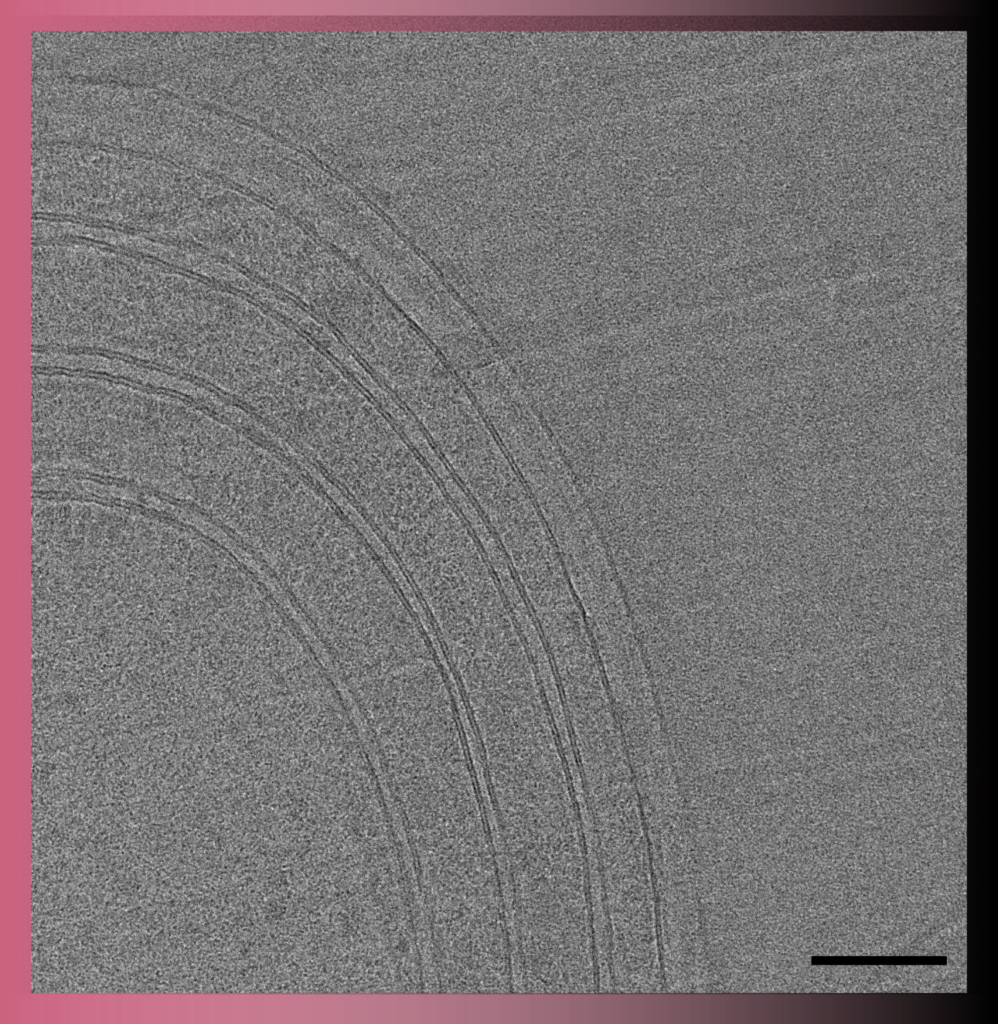
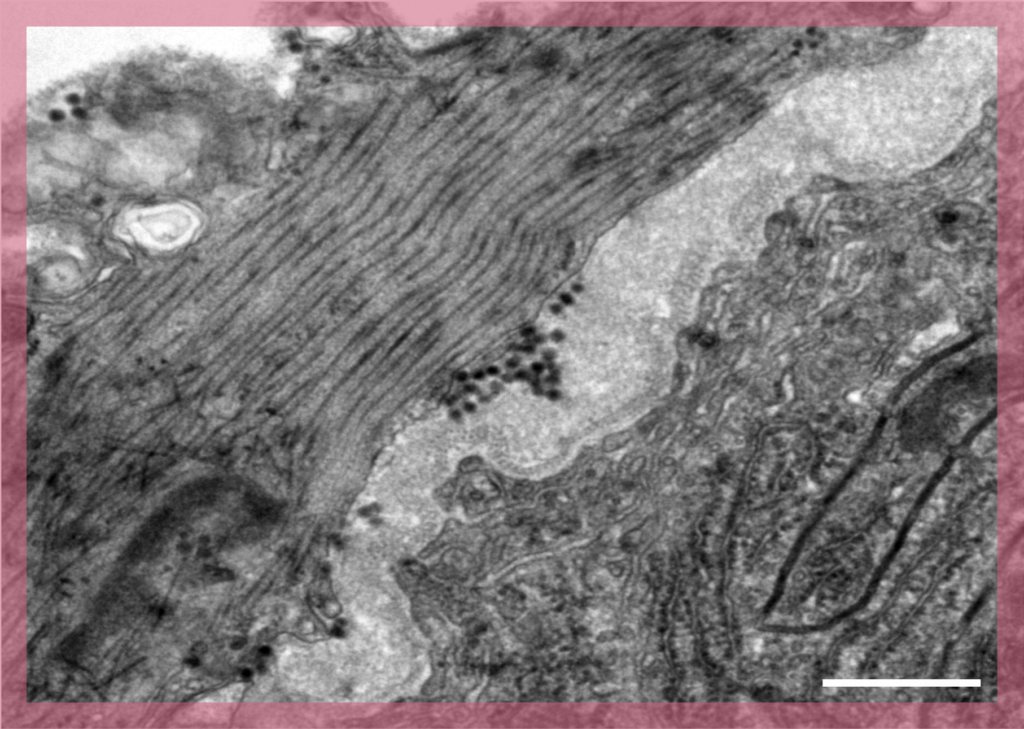
In Transmission Electron Microscopy (TEM) the image is formed from the interaction of the electrons with the sample as the beam is transmitted through the specimen. Transmission electron microscopes have the capability of imaging a biological sample at a significantly higher spatial resolution (ranging from nanometers to Angstroms) than any other microscope.
TEM allows imaging of small (such as viruses, proteins, liposomes, bacteria, archaea) or thin samples (such as the periphery of eukaryotic cells or sections of cells/tissue) with a size < 500 nm.
The UBI has a long term experience in imaging different samples using various TEM methods:
- Negative staining of isolated particles
- CryoEM automated acquisitions for single particle analysis
- Dual-axis tomography on plastic sections
- CryoET for single particles, cells and cryosections
- High Precision on grid CLEM and CryoCLEM
Sartori-Rupp A, Cordero Cervantes D, Pepe A, Gousset K, Delage E, Corroyer-Dulmont S, Schmitt C, Krijnse-Locker J. & C. Zurzolo, (2019). Correlative cryo-electron microscopy reveals the structure of TNTs in neuronal cells. Nat. Comm. 10, 342. Fédry J, Liu Y, Péhau-Arnaudet G, Pei J, Li W, Tortorici MA, Traincard F, Meola A, Bricogne G, Grishin NV, Snell WJ, Rey FA, & Krey T, (2017). The Ancient Gamete Fusogen HAP2 Is a Eukaryotic Class II Fusion Protein. Cell, 168(5), 904–915. Delaune A, Poupel O, Mallet A, Coic YM, Msadek T, Dubrac S. (2011) Peptidoglycan crosslinking relaxation plays an important role in Staphylococcus aureus WalKR-dependent cell viability. PLoS One. 6(2):e17054.
Scanning electron microscopy
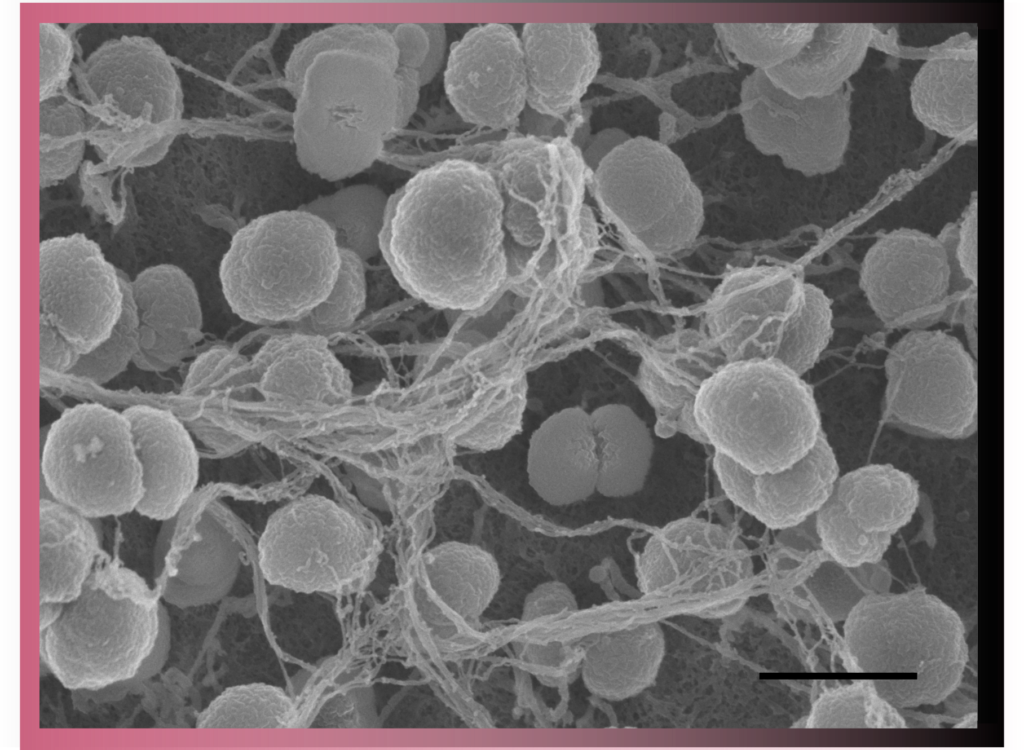
In Scanning Electron Microscopy (SEM) images are produced by scanning the surface of the sample with a focused beam of electrons. This is a routine technique used to characterise the surface of samples.
UBI has long-term experience with SEM and offers two scanning electron microscopes. These are equipped with secondary electron detectors and backscattered electron detectors for gold- immunolabelling experiments.
Grassart A, Malardé V, Gobaa S, Sartori-Rupp A, Kerns J, Karalis K, Marteyn B, Sansonetti P, Sauvonnet N. (2019). Bioengineered Human Organ-on-Chip Reveals Intestinal Microenvironment and Mechanical Forces Impacting Shigella Infection. Cell Host Microbe 26(3):435-444. Charles-Orszag A, Tsai FC, Bonazzi D, Manriquez V, Sachse M, Mallet A, Melican K, Staneva R et al., (2018) Adhesion to nanofibers drives cell membrane remodeling through one-dimension wetting. Nat Commun. 9(1):4450. Morello E, Mallet A, Konto-Ghiorghi Y et al. (2015). Evidence for the Sialylation of PilA, the PI-2a Pilus-Associated Adhesin of Streptococcus agalactiae Strain NEM316. PloS ONe, 10(9):e0138103.
Focused Ion Beam SEM
This imaging technology relies on combining serial milling and scanning of the resin bloc surface of the sample. The sequencial slicing and viewing steps therefor lead to the acquisition of large volumes in an automated way.
A Cryo-transfer module is installed for Cryo-FIB/SEM acquisitions. The production of thin cryo-lamella by FIB milling for subsequent cryoET collection and analysis is under development.
Quemin ER, Corroyer-Dulmont S, Baskaran A, Penard E, Gazi AD, Christo-Foroux E, Walther P, Abergel C, Krijnse-Locker J, (2019). Complex Membrane Remodeling during Virion Assembly of the 30,000-year-old Mollivirus Sibericum. J Virol. 2019 Jul 1; 93(13): e00388-19.
Fredlund J, Santos JC, Stevenin V, Weiner A, Latour-Lambert P, Rechav K, Mallet A, Krijnse-Locker J, Elbaum M, Enninga J, (2018). The entry of Salmonella in a distinct tight compartment revealed at high temporal and ultrastructural resolution. Cell Microbiol 20(4):e12816.
Correlative Light & Electron Microscopy
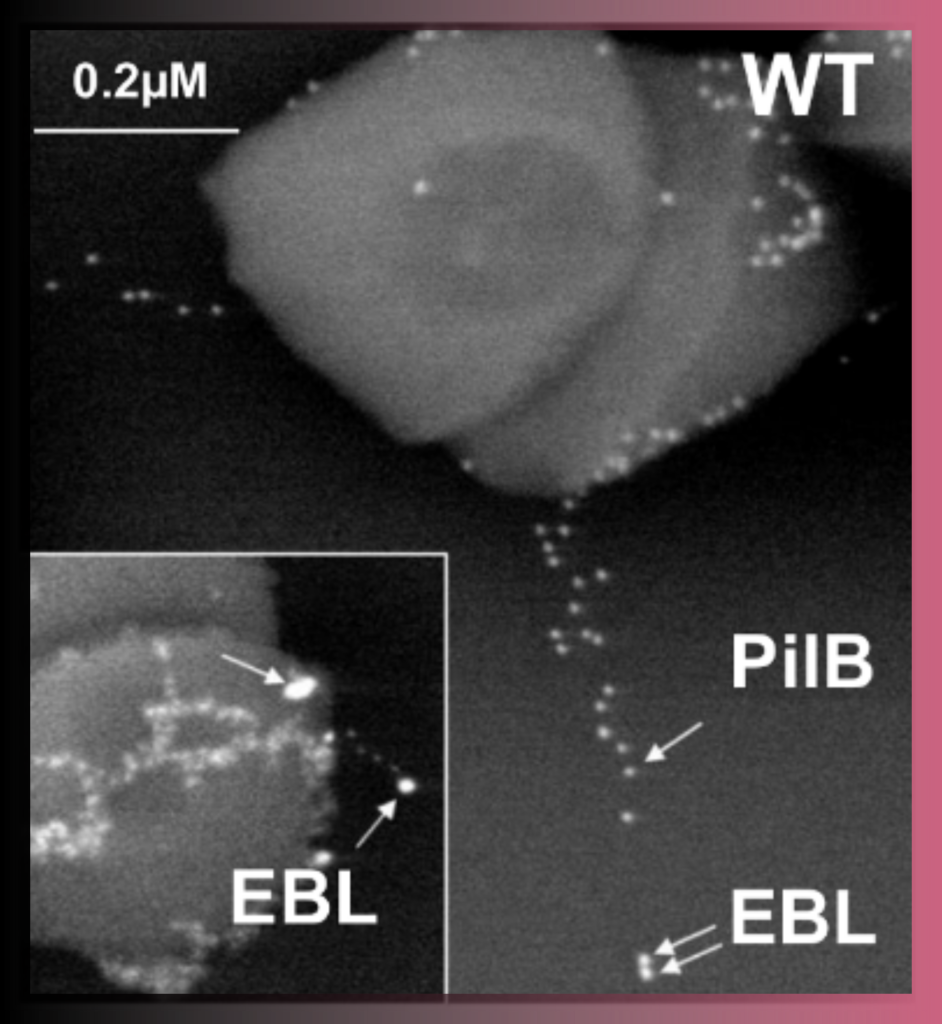
Correlative Light & Electron Microscopy (CLEM) approaches are powerful methods that combine the strengths of both light and electron microscopy. The functional and dynamic information coming from fluorescent light microscopy (FLM), such as widefield or confocal microscopy, can be directly linked to the higher resolution information provided by EM.
Various CLEM approaches are offered in UBI.
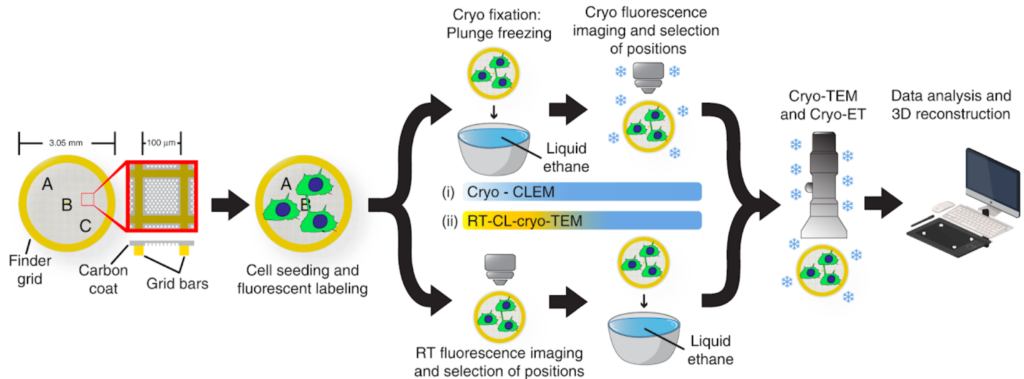
A) (High Precision) on-grid CLEM:
- Room Temperature CLEM
- Room Temperature Light Microscopy/CryoEM-CryoET
- Cryo-CLEM
B) Classical CLEM:
- CL-TEM
- CL-SEM – in combination with gold immunolabelling
- CL-FIB/SEM (Volume CLEM)
Sartori-Rupp A, Cordero Cervantes D, Pepe A, Gousset K, Delage E, Corroyer-Dulmont S, Schmitt C, Krijnse-Locker J. & C. Zurzolo, (2019). Correlative cryo-electron microscopy reveals the structure of TNTs in neuronal cells. Nat. Comm.10, 342. Quereda JJ, Sachse M, Balestrino D, Grenier T, Fredlund J, Danckaert A, Aulner N, Spenser S, Enninga J, Cossart P, Pizarro-Cerd
HT-Imaging for TEM 
Development of automated acquisition methods for TEM has been boosted mainly by academic efforts (UCSF Tomography, Leginon, SerialEM) in a relatively low cost. Since 2016 UBI is using SerialEM as an acquisition software package to automatize the collections on its Tecnai microscopes. To the time being UBI offers automated collections for:
-
Single Particles
-
Serial electron tomography (dual axis and CryoET)
-
High resolution mapping of entire cell profiles
-
On-grid CLEM approaches
The combination of serial sectioning together with the use of more complicated TEM tomography acquisition and data analysis schemes can provide the detailed information at high resolution needed for bigger volumes, even for entire infected cells. UBI has already a longstanding experience in serial sectioning, used until now mainly for classical CLEM approaches. As a next step UBI’s goal is to implement various acquisition protocols that can be used to search successfully for the areas of interest and enable specimen-specific image-analysis workflows for high-throughput data collection.